8GSU
 
 | | Crystal structure of human cardiac alpha actin (WT_ADP-Pi) in complex with fragmin F1 domain | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, Actin, ... | | Authors: | Iwasa, M, Oda, T, Takeda, S. | | Deposit date: | 2022-09-07 | | Release date: | 2023-03-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Mutagenic analysis of actin reveals the mechanism of His161 flipping that triggers ATP hydrolysis.
Front Cell Dev Biol, 11, 2023
|
|
8GT4
 
 | | Crystal structure of human cardiac alpha actin Q137A mutant (AMPPNP state) in complex with fragmin F1 domain | | Descriptor: | 1,2-ETHANEDIOL, Actin, alpha cardiac muscle 1, ... | | Authors: | Iwasa, M, Oda, T, Takeda, S. | | Deposit date: | 2022-09-07 | | Release date: | 2023-03-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Mutagenic analysis of actin reveals the mechanism of His161 flipping that triggers ATP hydrolysis.
Front Cell Dev Biol, 11, 2023
|
|
8GT1
 
 | | Crystal structure of human cardiac alpha actin A108G mutant (ADP-Pi state) in complex with fragmin F1 domain | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Iwasa, M, Oda, T, Takeda, S. | | Deposit date: | 2022-09-07 | | Release date: | 2023-03-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Mutagenic analysis of actin reveals the mechanism of His161 flipping that triggers ATP hydrolysis.
Front Cell Dev Biol, 11, 2023
|
|
8GSW
 
 | | Crystal structure of human cardiac alpha actin A108G mutant (AMPPNP state) in complex with fragmin F1 domain | | Descriptor: | 1,2-ETHANEDIOL, Actin, alpha cardiac muscle 1, ... | | Authors: | Iwasa, M, Oda, T, Takeda, S. | | Deposit date: | 2022-09-07 | | Release date: | 2023-03-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Mutagenic analysis of actin reveals the mechanism of His161 flipping that triggers ATP hydrolysis.
Front Cell Dev Biol, 11, 2023
|
|
8GT2
 
 | | Crystal structure of human cardiac alpha actin P109A mutant (AMPPNP state) in complex with fragmin F1 domain | | Descriptor: | 1,2-ETHANEDIOL, Actin, alpha cardiac muscle 1, ... | | Authors: | Iwasa, M, Oda, T, Takeda, S. | | Deposit date: | 2022-09-07 | | Release date: | 2023-03-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Mutagenic analysis of actin reveals the mechanism of His161 flipping that triggers ATP hydrolysis.
Front Cell Dev Biol, 11, 2023
|
|
8GT5
 
 | | Crystal structure of human cardiac alpha actin Q137A mutant (ADP-Pi state) in complex with fragmin F1 domain | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, Actin, ... | | Authors: | Iwasa, M, Oda, T, Takeda, S. | | Deposit date: | 2022-09-07 | | Release date: | 2023-03-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Mutagenic analysis of actin reveals the mechanism of His161 flipping that triggers ATP hydrolysis.
Front Cell Dev Biol, 11, 2023
|
|
8GT3
 
 | | Crystal structure of human cardiac alpha actin P109A mutant (ADP-Pi state) in complex with fragmin F1 domain | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, Actin, ... | | Authors: | Iwasa, M, Oda, T, Takeda, S. | | Deposit date: | 2022-09-07 | | Release date: | 2023-03-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Mutagenic analysis of actin reveals the mechanism of His161 flipping that triggers ATP hydrolysis.
Front Cell Dev Biol, 11, 2023
|
|
2ZGY
 
 | | PARM with GDP | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Plasmid segregation protein parM | | Authors: | Popp, D, Narita, A, Oda, T, Fujisawa, T, Matsuo, H, Nitanai, Y, Iwasa, M, Maeda, K, Onishi, H, Maeda, Y. | | Deposit date: | 2008-01-30 | | Release date: | 2008-02-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular structure of the ParM polymer and the mechanism leading to its nucleotide-driven dynamic instability
Embo J., 27, 2008
|
|
2ZHC
 
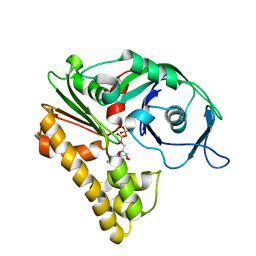 | | ParM filament | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Plasmid segregation protein parM | | Authors: | Popp, D, Narita, A, Oda, T, Fujisawa, T, Matsuo, H, Nitanai, Y, Iwasa, M, Maeda, K, Onishi, H, Maeda, Y. | | Deposit date: | 2008-02-04 | | Release date: | 2008-02-26 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (23 Å) | | Cite: | Molecular structure of the ParM polymer and the mechanism leading to its nucleotide-driven dynamic instability
Embo J., 27, 2008
|
|
2ZWH
 
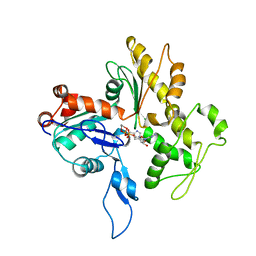 | | Model for the F-actin structure | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin, alpha skeletal muscle, ... | | Authors: | Oda, T, Iwasa, M, Aihara, T, Maeda, Y, Narita, A. | | Deposit date: | 2008-12-05 | | Release date: | 2009-01-20 | | Last modified: | 2017-10-11 | | Method: | FIBER DIFFRACTION (3.3 Å) | | Cite: | The nature of the globular- to fibrous-actin transition.
Nature, 457, 2009
|
|
2ZGZ
 
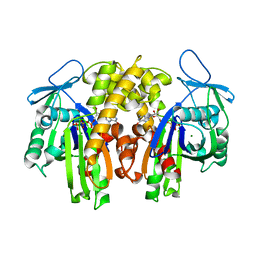 | | PARM with GMPPNP | | Descriptor: | MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, Plasmid segregation protein parM | | Authors: | Popp, D, Narita, A, Oda, T, Fujisawa, T, Matsuo, H, Nitanai, Y, Iwasa, M, Maeda, K, Onishi, H, Maeda, Y. | | Deposit date: | 2008-01-30 | | Release date: | 2008-02-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Molecular structure of the ParM polymer and the mechanism leading to its nucleotide-driven dynamic instability
Embo J., 27, 2008
|
|
3BIS
 
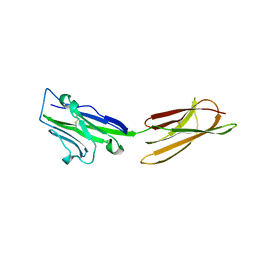 | | Crystal Structure of the PD-L1 | | Descriptor: | Programmed cell death 1 ligand 1 | | Authors: | Lin, D.Y, Tanaka, Y, Iwasaki, M, Gittis, A.G, Su, H.P, Mikami, B, Okazaki, T, Honjo, T, Minato, N, Garboczi, D.N. | | Deposit date: | 2007-11-30 | | Release date: | 2008-02-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | The PD-1/PD-L1 complex resembles the antigen-binding Fv domains of antibodies and T cell receptors.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3BIK
 
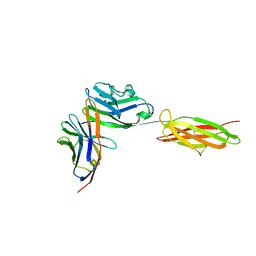 | | Crystal Structure of the PD-1/PD-L1 Complex | | Descriptor: | GLYCEROL, Programmed cell death 1 ligand 1, Programmed cell death protein 1 | | Authors: | Lin, D.Y, Tanaka, Y, Iwasaki, M, Gittis, A.G, Su, H.P, Mikami, B, Okazaki, T, Honjo, T, Minato, N, Garboczi, D.N. | | Deposit date: | 2007-11-30 | | Release date: | 2008-02-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | The PD-1/PD-L1 complex resembles the antigen-binding Fv domains of antibodies and T cell receptors.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
7W51
 
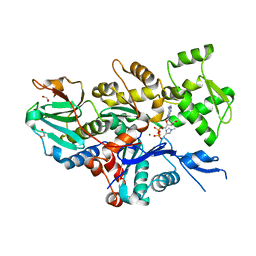 | |
7W4Z
 
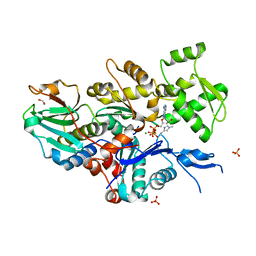 | |
7W50
 
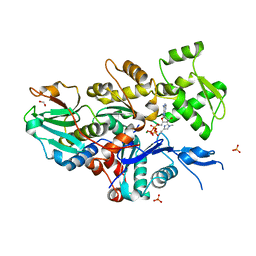 | |
7W52
 
 | |
7YNE
 
 | |
3I9T
 
 | | Crystal structure of the rat heme oxygenase (HO-1) in complex with heme binding dithiothreitol (DTT) | | Descriptor: | (2S,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, Heme oxygenase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Matsui, T, Unno, M, Ikeda-Saito, M. | | Deposit date: | 2009-07-13 | | Release date: | 2010-05-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Dioxygen activation for the self-degradation of heme: reaction mechanism and regulation of heme oxygenase.
Inorg.Chem., 49, 2010
|
|
3I9U
 
 | | Crystal structure of the rat heme oxygenase (HO-1) in complex with heme binding dithioerythritol (DTE) | | Descriptor: | (2R,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, Heme oxygenase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Matsui, T, Unno, M, Ikeda-Saito, M. | | Deposit date: | 2009-07-13 | | Release date: | 2010-05-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Dioxygen activation for the self-degradation of heme: reaction mechanism and regulation of heme oxygenase.
Inorg.Chem., 49, 2010
|
|
3I8R
 
 | | Crystal structure of the heme oxygenase from Corynebacterium diphtheriae (HmuO) in complex with heme binding ditiothreitol (DTT) | | Descriptor: | (2S,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, Heme oxygenase, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Matsui, T, Unno, M, Ikeda-Saito, M. | | Deposit date: | 2009-07-10 | | Release date: | 2010-05-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Dioxygen activation for the self-degradation of heme: reaction mechanism and regulation of heme oxygenase.
Inorg.Chem., 49, 2010
|
|
