1MCT
 
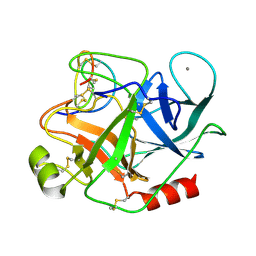 | | THE REFINED 1.6 ANGSTROMS RESOLUTION CRYSTAL STRUCTURE OF THE COMPLEX FORMED BETWEEN PORCINE BETA-TRYPSIN AND MCTI-A, A TRYPSIN INHIBITOR OF SQUASH FAMILY | | Descriptor: | BETA-TRYPSIN, CALCIUM ION, TRYPSIN INHIBITOR A | | Authors: | Huang, Q, Liu, S, Tang, Y. | | Deposit date: | 1992-10-24 | | Release date: | 1994-01-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Refined 1.6 A resolution crystal structure of the complex formed between porcine beta-trypsin and MCTI-A, a trypsin inhibitor of the squash family. Detailed comparison with bovine beta-trypsin and its complex.
J.Mol.Biol., 229, 1993
|
|
1MRJ
 
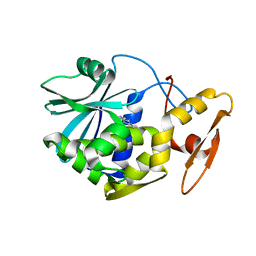 | | STUDIES ON CRYSTAL STRUCTURES ACTIVE CENTER GEOMETRY AND DEPURINE MECHANISM OF TWO RIBOSOME-INACTIVATING PROTEINS | | Descriptor: | ADENOSINE, ALPHA-TRICHOSANTHIN | | Authors: | Huang, Q, Liu, S, Tang, Y, Jin, S, Wang, Y. | | Deposit date: | 1994-07-01 | | Release date: | 1995-02-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Studies on crystal structures, active-centre geometry and depurinating mechanism of two ribosome-inactivating proteins.
Biochem.J., 309, 1995
|
|
5D3O
 
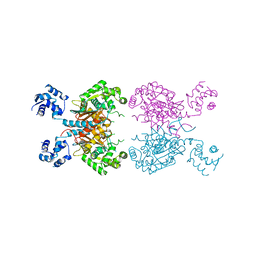 | |
8DZT
 
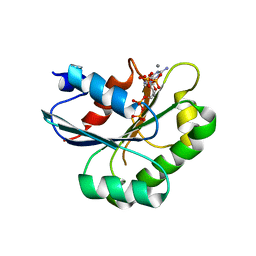 | | Crystal structure of human Sar1aH79G mutant | | Descriptor: | CALCIUM ION, GTP-binding protein SAR1a, GUANOSINE-5',3'-TETRAPHOSPHATE, ... | | Authors: | Huang, Q. | | Deposit date: | 2022-08-08 | | Release date: | 2022-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8E0A
 
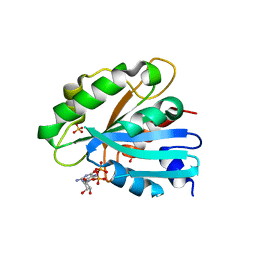 | | Crystal structure of human Sar1b | | Descriptor: | GTP-binding protein SAR1b, GUANOSINE-5'-DIPHOSPHATE, SULFATE ION | | Authors: | Huang, Q. | | Deposit date: | 2022-08-08 | | Release date: | 2022-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.797 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8E0D
 
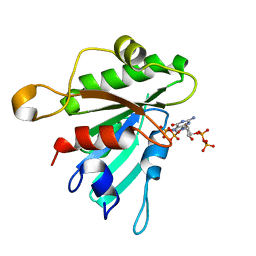 | | Crystal structure of human Sar1bE140D | | Descriptor: | GTP-binding protein SAR1b, GUANOSINE-5',3'-TETRAPHOSPHATE | | Authors: | Huang, Q. | | Deposit date: | 2022-08-08 | | Release date: | 2022-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.981 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8E0H
 
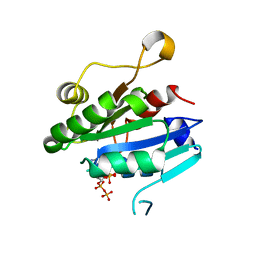 | | Crystal structure of human Sar1aD104/D140A double mutant | | Descriptor: | GTP-binding protein SAR1a, GUANOSINE-5',3'-TETRAPHOSPHATE | | Authors: | Huang, Q. | | Deposit date: | 2022-08-09 | | Release date: | 2022-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8DZO
 
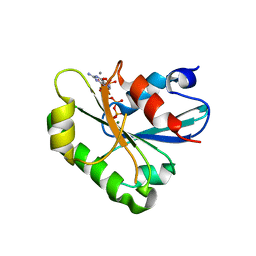 | | Crystal structure of human Sar1T39N mutant | | Descriptor: | CALCIUM ION, GTP-binding protein SAR1a, GUANOSINE-5',3'-TETRAPHOSPHATE, ... | | Authors: | Huang, Q. | | Deposit date: | 2022-08-08 | | Release date: | 2022-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8E0B
 
 | | Crystal structure of human Sar1bT39N | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, SULFATE ION, Sar1bT39N | | Authors: | Huang, Q. | | Deposit date: | 2022-08-08 | | Release date: | 2023-08-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.212 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8E0C
 
 | | Crystal structure of human Sar1bH79G | | Descriptor: | GTP-binding protein SAR1b, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION | | Authors: | Huang, Q. | | Deposit date: | 2022-08-08 | | Release date: | 2023-08-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.993 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8DZN
 
 | | Crystal structure of human Sar1a in complex with GDP | | Descriptor: | GTP-binding protein SAR1a, GUANOSINE-5'-DIPHOSPHATE, SULFATE ION | | Authors: | Huang, Q. | | Deposit date: | 2022-08-08 | | Release date: | 2023-08-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.107 Å) | | Cite: | The alarmone ppGpp selectively inhibits the isoform A of the human small GTPase Sar1.
Proteins, 91, 2023
|
|
8DZM
 
 | |
2FM5
 
 | | Crystal structure of PDE4D2 in complex with inhibitor L-869299 | | Descriptor: | (R)-3-(2-(3-CYCLOPROPOXY-4-(DIFLUOROMETHOXY)PHENYL)-2-(5-(1,1,1,3,3,3-HEXAFLUORO-2-HYDROXYPROPAN-2-YL)THIAZOL-2-YL)ETHYL)PYRIDINE 1-OXIDE, MAGNESIUM ION, ZINC ION, ... | | Authors: | Huai, Q, Sun, Y, Wang, H, Macdonald, D, Aspiotis, R, Robinson, H, Huang, Z, Ke, H. | | Deposit date: | 2006-01-07 | | Release date: | 2006-03-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Enantiomer Discrimination Illustrated by the High Resolution Crystal Structures of Type 4 Phosphodiesterase
J.Med.Chem., 49, 2006
|
|
2FM0
 
 | | Crystal structure of PDE4D in complex with L-869298 | | Descriptor: | (S)-3-(2-(3-CYCLOPROPOXY-4-(DIFLUOROMETHOXY)PHENYL)-2-(5-(1,1,1,3,3,3-HEXAFLUORO-2-HYDROXYPROPAN-2-YL)THIAZOL-2-YL)ETHY L)PYRIDINE 1-OXIDE, MAGNESIUM ION, ZINC ION, ... | | Authors: | Huai, Q, Sun, Y, Wang, H, Macdonald, D, Aspiotis, R, Robinson, H, Huang, Z, Ke, H. | | Deposit date: | 2006-01-06 | | Release date: | 2006-03-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Enantiomer Discrimination Illustrated by the High Resolution Crystal Structures of Type 4 Phosphodiesterase
J.Med.Chem., 49, 2006
|
|
3BI5
 
 | |
3BI4
 
 | | Crystal structures of fms1 in complex with its inhibitors | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, N-[(1E,2Z)-but-2-en-1-ylidene]-N'-[(2E)-but-2-en-1-ylidene]butane-1,4-diamine, Polyamine oxidase FMS1 | | Authors: | Huang, Q, Hao, Q. | | Deposit date: | 2007-11-29 | | Release date: | 2008-01-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of fms1 in complex with its inhibitors
To be Published
|
|
3BI2
 
 | |
3BNM
 
 | | Crystal structure of polyamine oxidase FMS1 from Saccharomyces cerevisiae in complex with bis-(3R,3'R)-methylated spermine | | Descriptor: | (3R,3'R)-N~1~,N~1~'-butane-1,4-diyldibutane-1,3-diamine, FLAVIN-ADENINE DINUCLEOTIDE, Polyamine oxidase FMS1 | | Authors: | Huang, Q, Hao, Q. | | Deposit date: | 2007-12-14 | | Release date: | 2008-01-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural basis of the substrate stereospecificity of FMS1.
To Be Published
|
|
3BNU
 
 | | Crystal structure of polyamine oxidase FMS1 from Saccharomyces cerevisiae in complex with bis-(3S,3'S)-methylated spermine | | Descriptor: | (3S,3'S)-N~1~,N~1~'-butane-1,4-diyldibutane-1,3-diamine, FLAVIN-ADENINE DINUCLEOTIDE, Polyamine oxidase FMS1 | | Authors: | Huang, Q, Hao, Q. | | Deposit date: | 2007-12-14 | | Release date: | 2008-01-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural basis of the substrate stereospecificity of FMS1.
To Be Published
|
|
1Z6L
 
 | |
1RSG
 
 | |
1YY5
 
 | |
1XPQ
 | | Crystal structure of fms1, a polyamine oxidase from yeast | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Polyamine oxidase FMS1, SPERMINE | | Authors: | Huang, Q, Liu, Q, Hao, Q. | | Deposit date: | 2004-10-09 | | Release date: | 2005-04-26 | | Last modified: | 2021-12-08 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Crystal structures of Fms1 and its complex with spermine reveal substrate specificity.
J.Mol.Biol., 348, 2005
|
|
6UJG
 | | Crystal structure of human GAC in complex with inhibitor UPGL00012 | | Descriptor: | Glutaminase kidney isoform, mitochondrial, N-{5-[(3S)-3-{[5-(acetylamino)-1,3,4-thiadiazol-2-yl]amino}pyrrolidin-1-yl]-1,3,4-thiadiazol-2-yl}acetamide | | Authors: | Huang, Q, Cerione, R.A. | | Deposit date: | 2019-10-03 | | Release date: | 2020-10-07 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of human GAC in complex with inhibitor UPGL00012
To Be Published
|
|
6UME
 
 | | Crystal structure of human GAC in complex with inhibitor UPGL00012 | | Descriptor: | 2-(pyridin-3-yl)-N-(5-{4-[(5-{[(pyridin-3-yl)acetyl]amino}-1,3,4-thiadiazol-2-yl)amino]piperidin-1-yl}-1,3,4-thiadiazol-2-yl)acetamide, Glutaminase kidney isoform, mitochondrial | | Authors: | Huang, Q, Cerione, R.A. | | Deposit date: | 2019-10-09 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of human GAC in complex with inhibitor UPGL00012
To Be Published
|
|
