2FLF
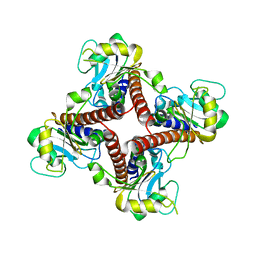 
 | | Crystal structure of l-fuculose-1-phosphate aldolase from Thermus Thermophilus HB8 | | Descriptor: | fuculose-1-phosphate aldolase | | Authors: | Jeyakanthan, J, Yokoyama, S, Shiro, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-01-06 | | Release date: | 2007-01-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Purification, crystallization and preliminary X-ray crystallographic study of the L-fuculose-1-phosphate aldolase (FucA) from Thermus thermophilus HB8
Acta Crystallogr.,Sect.F, 61, 2005
|
|
2DY8
 
 | |
2CZY
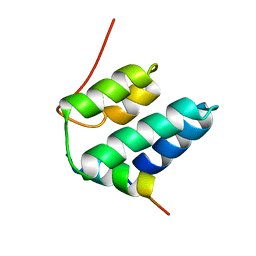 
 | | Solution structure of the NRSF/REST-mSin3B PAH1 complex | | Descriptor: | Paired amphipathic helix protein Sin3b, transcription factor REST (version 3) | | Authors: | Nomura, M, Uda-Tochio, H, Murai, K, Mori, N, Nishimura, Y. | | Deposit date: | 2005-07-20 | | Release date: | 2005-12-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The Neural Repressor NRSF/REST Binds the PAH1 Domain of the Sin3 Corepressor by Using its Distinct Short Hydrophobic Helix
J.Mol.Biol., 354, 2005
|
|
2DPT
 
 | |
2DPS
 
 | |
2E8J
 
 | |
2ERB
 
 | | AgamOBP1, and odorant binding protein from Anopheles gambiae complexed with PEG | | Descriptor: | 2,5,8,11,14,17,20,23,26,29,32,35,38,41,44,47,50,53,56,59,62,65,68,71,74,77,80-HEPTACOSAOXADOOCTACONTAN-82-OL, MAGNESIUM ION, odorant binding protein | | Authors: | Wogulis, M, Morgan, T, Ishida, Y, Leal, W.S, Wilson, D.K. | | Deposit date: | 2005-10-24 | | Release date: | 2005-12-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The crystal structure of an odorant binding protein from Anopheles gambiae: Evidence for a common ligand release mechanism.
Biochem.Biophys.Res.Commun., 339, 2006
|
|
2CMM
 
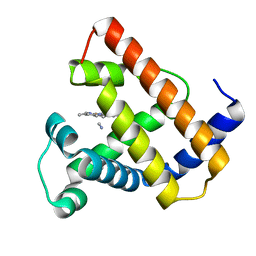 | | STRUCTURAL ANALYSIS OF THE MYOGLOBIN RECONSTITUTED WITH IRON PORPHINE | | Descriptor: | CYANIDE ION, MYOGLOBIN, PORPHYRIN FE(III) | | Authors: | Sato, T, Tanaka, N, Moriyama, H, Igarashi, N, Neya, S, Funasaki, N, Iizuka, T, Shiro, Y. | | Deposit date: | 1993-12-24 | | Release date: | 1994-01-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural analysis of the myoglobin reconstituted with iron porphine.
J.Biol.Chem., 268, 1993
|
|
2FKO
 
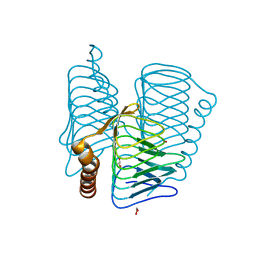 | | Structure of PH1591 from Pyrococcus horikoshii OT3 | | Descriptor: | 1,2-ETHANEDIOL, 173aa long hypothetical ferripyochelin binding protein, ZINC ION | | Authors: | Jeyakanthan, J, Tahirov, T.H, Yokoyama, S, Shiro, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-01-05 | | Release date: | 2007-01-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Observation of a calcium-binding site in the gamma-class carbonic anhydrase from Pyrococcus horikoshii.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
2F7L
 
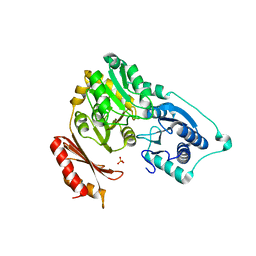 | | Crystal structure of Sulfolobus tokodaii phosphomannomutase/phosphoglucomutase | | Descriptor: | 455aa long hypothetical phospho-sugar mutase, SULFATE ION | | Authors: | Kawamura, T, Sakai, N, Akutsu, J, Zhang, Z, Watanabe, N, Kawarabayashi, Y, Tanaka, I. | | Deposit date: | 2005-12-01 | | Release date: | 2006-12-05 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of Sulfolobus tokodaii phosphomannomutase/phosphoglucomutase
To be Published
|
|
2D21
 
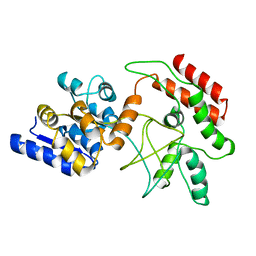 | | NMR Structure of stereo-array isotope labelled (SAIL) maltodextrin-binding protein (MBP) | | Descriptor: | Maltose-binding periplasmic protein | | Authors: | Kainosho, M, Torizawa, T, Iwashita, Y, Terauchi, T, Ono, A.M, Guntert, P. | | Deposit date: | 2005-09-02 | | Release date: | 2006-03-07 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Optimal isotope labelling for NMR protein structure determinations.
Nature, 440, 2006
|
|
3K9X
 
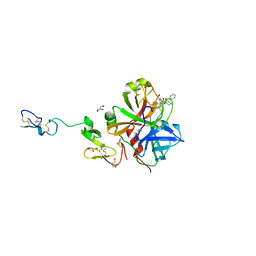 | | X-ray crystal structure of human fxa in complex with (S)-N-((2-METHYLBENZOFURAN-5-YLAMINO)(2-OXO-1-(2-OXO-2- (PYRROLIDIN-1-YL)ETHYL)AZEPAN-3- YLAMINO)METHYLENE)NICOTINAMIDE | | Descriptor: | CALCIUM ION, GLYCEROL, N-{N'-(2-methyl-1-benzofuran-5-yl)-N-[(3S)-2-oxo-1-(2-oxo-2-pyrrolidin-1-ylethyl)azepan-3-yl]carbamimidoyl}pyridine-3-carboxamide, ... | | Authors: | Klei, H.E, Kish, K, Ghosh, K, Rushith, A. | | Deposit date: | 2009-10-16 | | Release date: | 2009-12-15 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Aroylguanidine-based factor Xa inhibitors: the discovery of BMS-344577
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3HPT
 
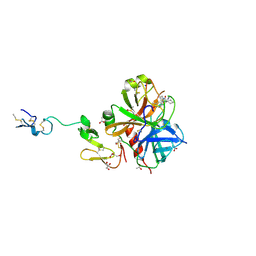 | | Crystal structure of human FxA in complex with (S)-2-cyano-1-(2-methylbenzofuran-5-yl)-3-(2-oxo-1-(2-oxo-2-(pyrrolidin-1-yl)ethyl)azepan-3-yl)guanidine | | Descriptor: | 1-cyano-2-(2-methyl-1-benzofuran-5-yl)-3-[(3S)-2-oxo-1-(2-oxo-2-pyrrolidin-1-ylethyl)azepan-3-yl]guanidine, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ACETATE ION, ... | | Authors: | Klei, H.E, Ghosh, K, Rushith, A, Kish, K. | | Deposit date: | 2009-06-04 | | Release date: | 2009-11-17 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Cyanoguanidine-based lactam derivatives as a novel class of orally bioavailable factor Xa inhibitors.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
7NAK
 
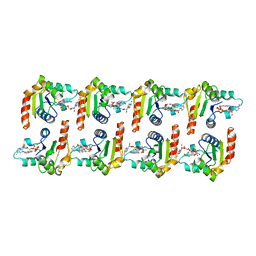 | | Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (TIR:1AD) | | Descriptor: | NAD(+) hydrolase SARM1, [[(2~{R},3~{S},4~{R},5~{R})-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] [(2~{R},3~{S},4~{R},5~{R})-5-(5-iodanylisoquinolin-2-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methyl hydrogen phosphate | | Authors: | Kerry, P.S, Nanson, J.D, Adams, S, Cunnea, K, Bosanac, T, Kobe, B, Hughes, R.O, Ve, T. | | Deposit date: | 2021-06-21 | | Release date: | 2022-03-23 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural basis of SARM1 activation, substrate recognition, and inhibition by small molecules.
Mol.Cell, 82, 2022
|
|
7NAL
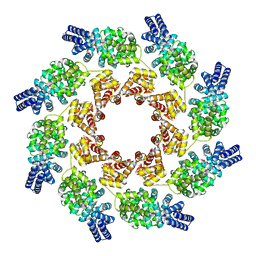 
 | | Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (ARM and SAM domains) | | Descriptor: | BETA-NICOTINAMIDE RIBOSE MONOPHOSPHATE, NAD(+) hydrolase SARM1 | | Authors: | Kerry, P.S, Nanson, J.D, Adams, S, Cunnea, K, Bosanac, T, Kobe, B, Hughes, R.O, Ve, T. | | Deposit date: | 2021-06-21 | | Release date: | 2022-03-23 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural basis of SARM1 activation, substrate recognition, and inhibition by small molecules.
Mol.Cell, 82, 2022
|
|
7T79
 
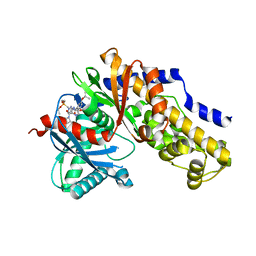 | |
7T78
 
 | |
6LKX
 
 | | The structure of PRRSV helicase | | Descriptor: | CITRIC ACID, GLYCEROL, RNA-dependent RNA polymerase, ... | | Authors: | Shi, Y.J, Tong, X.H, Peng, G.Q. | | Deposit date: | 2019-12-20 | | Release date: | 2020-05-27 | | Last modified: | 2021-06-09 | | Method: | X-RAY DIFFRACTION (2.998 Å) | | Cite: | Structural Characterization of the Helicase nsp10 Encoded by Porcine Reproductive and Respiratory Syndrome Virus.
J.Virol., 94, 2020
|
|
3SW2
 
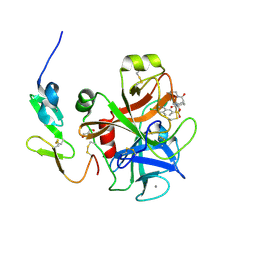 | | X-ray crystal structure of human FXA in complex with 6-chloro-N-((3S)-2-oxo-1-(2-oxo-2-((5S)-8-oxo-5,6-dihydro-1H-1,5-methanopyrido[1,2-a][1,5]diazocin-3(2H,4H,8H)-yl)ethyl)piperidin-3-yl)naphthalene-2-sulfonamide | | Descriptor: | 6-chloro-N-((3S)-2-oxo-1-(2-oxo-2-((5S)-8-oxo-5,6-dihydro-1H-1,5-methanopyrido[1,2-a][1,5]diazocin-3(2H,4H,8H)-yl)ethyl)piperidin-3-yl)naphthalene-2-sulfonamide, CALCIUM ION, Coagulation factor X, ... | | Authors: | Klei, H.E. | | Deposit date: | 2011-07-13 | | Release date: | 2011-11-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Arylsulfonamidopiperidone derivatives as a novel class of factor Xa inhibitors.
Bioorg.Med.Chem.Lett., 21, 2011
|
|
3ENS
 
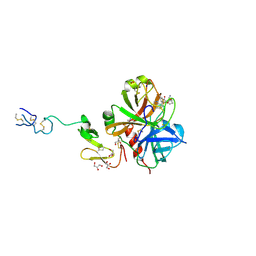 | |
2N3T
 
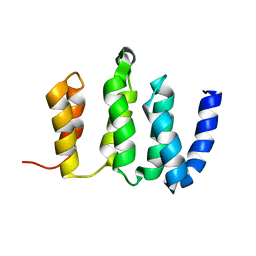 | |
2N3W
 
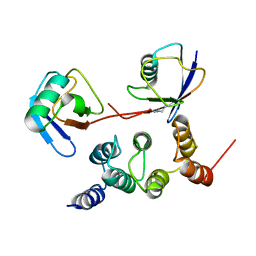 | |
2N3U
 
 | |
2N3V
 
 | |
6KFW
 
 | |
