8PDK
 
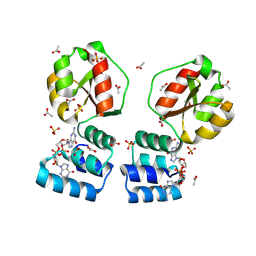 | | X-ray structure of the Thermus thermophilus PilF-GSPIIB domain in the c-di-GMP bound state | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), ACETATE ION, SULFATE ION, ... | | Authors: | Neissner, K, Woehnert, J. | | Deposit date: | 2023-06-12 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | X-ray structure of the PilF-GSPIIB domain in the c-di-GMP bound state
To Be Published
|
|
8PFA
 
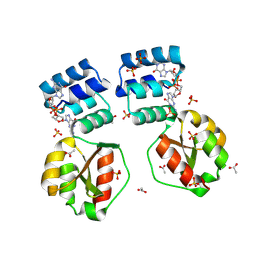 | | X-ray structure of the Thermus thermophilus K167R mutant of the PilF-GSPIIB domain in the c-di-GMP bound state | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), ACETATE ION, SULFATE ION, ... | | Authors: | Neissner, K, Woehnert, J. | | Deposit date: | 2023-06-15 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray structure of the Thermus thermophilus K167R mutant of the PilF-GSPIIB domain in the c-di-GMP bound state
To Be Published
|
|
8PQU
 
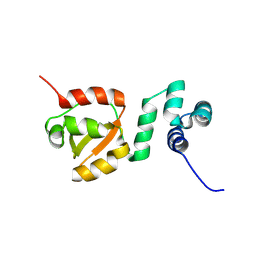 | |
8PKZ
 
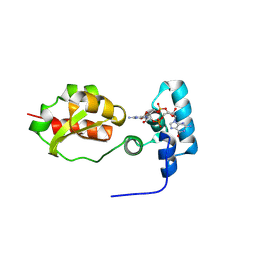 | |
8A2N
 | | Structure of crocagin biosynthetic protein CgnD | | Descriptor: | CgnD, SULFATE ION | | Authors: | Adam, S, Koehnke, J. | | Deposit date: | 2022-06-06 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Unusual peptide-binding proteins guide pyrroloindoline alkaloid formation in crocagin biosynthesis.
Nat.Chem., 15, 2023
|
|
8R1B
 
 | |
8AHD
 
 | |
8AGY
 
 | |
2X5N
 
 | | Crystal Structure of the SpRpn10 VWA domain | | Descriptor: | 26S PROTEASOME REGULATORY SUBUNIT RPN10, SULFATE ION | | Authors: | Riedinger, C, Boehringer, J, Trempe, J.-F, Lowe, E.D, Brown, N.R, Gehring, K, Noble, M.E.M, Gordon, C, Endicott, J.A. | | Deposit date: | 2010-02-10 | | Release date: | 2010-08-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The Structure of Rpn10 and its Interactions with Polyubiquitin Chains and the Proteasome Subunit Rpn12.
J.Biol.Chem., 285, 2010
|
|
5M9N
 
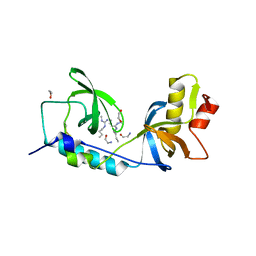 | | Crystal structure of human TDRD1 extended Tudor domain in complex with a symmetrically dimethylated E2F peptide | | Descriptor: | 1,2-ETHANEDIOL, E2F peptide, N3, ... | | Authors: | Tallant, C, Savitsky, P, Moehlenbrink, J, Chan, C, Nunez-Alonso, G, Newman, J.A, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Fedorov, O, La Thangue, N.B, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2016-11-01 | | Release date: | 2016-11-16 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of human TDRD1 extended Tudor domain in complex with a symmetrically
dimethylated E2F peptide
To Be Published
|
|
5M9O
 
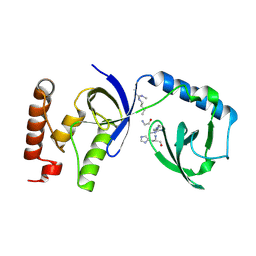 | | Crystal structure of human SND1 extended Tudor domain in complex with a symmetrically dimethylated E2F peptide | | Descriptor: | E2F peptide, Staphylococcal nuclease domain-containing protein 1 | | Authors: | Tallant, C, Savitsky, P, Moehlenbrink, J, Chan, C, Nunez-Alonso, G, Siejka, P, Sorrell, F.J, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Fedorov, O, La Thangue, N.B, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2016-11-01 | | Release date: | 2016-12-07 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crystal structure of human SND1 extended Tudor domain in complex with a symmetrically
dimethylated E2F peptide
To Be Published
|
|
1EUB
 
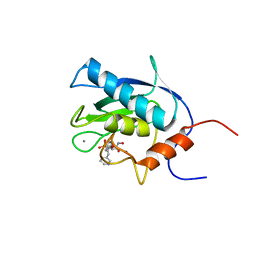 | | SOLUTION STRUCTURE OF THE CATALYTIC DOMAIN OF HUMAN COLLAGENASE-3 (MMP-13) COMPLEXED TO A POTENT NON-PEPTIDIC SULFONAMIDE INHIBITOR | | Descriptor: | 1-METHYLOXY-4-SULFONE-BENZENE, 3-METHYLPYRIDINE, CALCIUM ION, ... | | Authors: | Zhang, X, Gonnella, N.C, Koehn, J, Pathak, N, Ganu, V, Melton, R, Parker, D, Hu, S.I, Nam, K.Y. | | Deposit date: | 2000-04-14 | | Release date: | 2001-04-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the catalytic domain of human collagenase-3 (MMP-13) complexed to a potent non-peptidic sulfonamide inhibitor: binding comparison with stromelysin-1 and collagenase-1.
J.Mol.Biol., 301, 2000
|
|
6I86
 
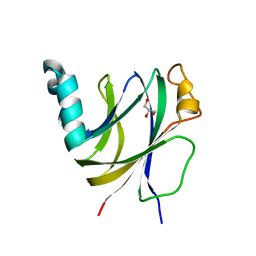 | | Crocagin biosynthetic gene J | | Descriptor: | GLYCEROL, Uncharacterized protein | | Authors: | Adam, S, Koehnke, J. | | Deposit date: | 2018-11-19 | | Release date: | 2019-03-13 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structure of CgnJ, a domain of unknown function protein from the crocagin gene cluster.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
2JQ7
 
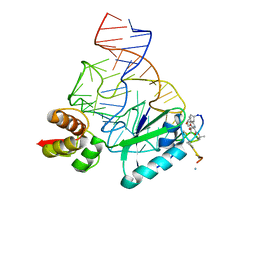 | | Model for thiostrepton binding to the ribosomal L11-RNA | | Descriptor: | 50S RIBOSOMAL PROTEIN L11, RIBOSOMAL RNA, THIOSTREPTON | | Authors: | Jonker, H.R.A, Ilin, S, Grimm, S.K, Woehnert, J, Schwalbe, H. | | Deposit date: | 2007-05-30 | | Release date: | 2007-07-03 | | Last modified: | 2021-08-18 | | Method: | SOLUTION NMR | | Cite: | L11 Domain Rearrangement Upon Binding to RNA and Thiostrepton Studied by NMR Spectroscopy
Nucleic Acids Res., 35, 2007
|
|
8OO2
 
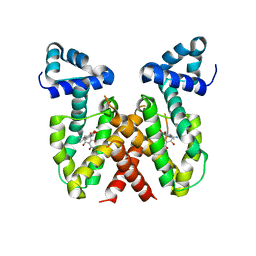 | | ChdA complex with amido-chelocardin | | Descriptor: | 2-carboxamido-2-deacetyl-chelocardin, MAGNESIUM ION, Putative transcriptional regulator | | Authors: | Koehnke, J, Sikandar, A. | | Deposit date: | 2023-04-04 | | Release date: | 2023-08-02 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Revision of the Absolute Configurations of Chelocardin and Amidochelocardin.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8CH5
 | |
3MW2
 | | Crystal structure of beta-neurexin 1 with the splice insert 4 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-D-mannopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Neurexin-1-alpha, PHOSPHATE ION | | Authors: | Jin, X, Shapiro, L. | | Deposit date: | 2010-05-05 | | Release date: | 2010-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Splice Form Dependence of beta-Neurexin/Neuroligin Binding Interactions.
Neuron, 67, 2010
|
|
3MW4
 
 | | Crystal structure of beta-neurexin 3 without the splice insert 4 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-D-mannopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Neurexin-2-beta, ... | | Authors: | Jin, X, Shapiro, L. | | Deposit date: | 2010-05-05 | | Release date: | 2010-07-28 | | Last modified: | 2021-10-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Splice Form Dependence of beta-Neurexin/Neuroligin Binding Interactions.
Neuron, 67, 2010
|
|
3MW3
 
 | | Crystal structure of beta-neurexin 2 with the splice insert 4 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-D-mannopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Neurexin-2-beta | | Authors: | Jin, X, Shapiro, L. | | Deposit date: | 2010-05-05 | | Release date: | 2010-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Splice Form Dependence of beta-Neurexin/Neuroligin Binding Interactions.
Neuron, 67, 2010
|
|
8AUR
 
 | | Cryo-EM structure of a TasA fibre | | Descriptor: | Major biofilm matrix component | | Authors: | Boehning, J, Bharat, T.A.M. | | Deposit date: | 2022-08-25 | | Release date: | 2022-11-30 | | Last modified: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (3.47 Å) | | Cite: | Donor-strand exchange drives assembly of the TasA scaffold in Bacillus subtilis biofilms.
Nat Commun, 13, 2022
|
|
8CIO
 
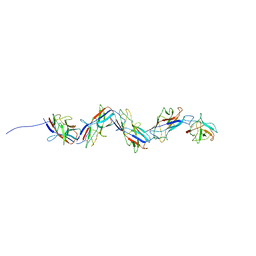 | |
6F8B
 
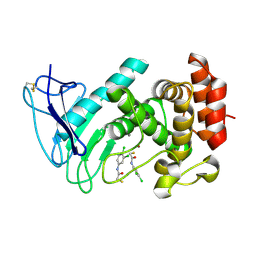 | | LasB bound to thiol based inhibitor | | Descriptor: | CALCIUM ION, Elastase, ZINC ION, ... | | Authors: | Koehnke, J, Sikandar, A. | | Deposit date: | 2017-12-12 | | Release date: | 2018-03-28 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Binding Mode Characterization and Early in Vivo Evaluation of Fragment-Like Thiols as Inhibitors of the Virulence Factor LasB from Pseudomonas aeruginosa.
ACS Infect Dis, 4, 2018
|
|
6FZX
 
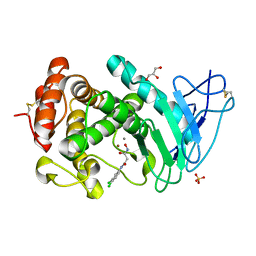 | | LasB, hydroxymate Inhibitor Complex | | Descriptor: | CALCIUM ION, GLYCEROL, Keratinase KP2, ... | | Authors: | Koehnke, J, Sikandar, A. | | Deposit date: | 2018-03-15 | | Release date: | 2018-09-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Tackling Pseudomonas aeruginosa Virulence by a Hydroxamic Acid-Based LasB Inhibitor.
ACS Chem. Biol., 13, 2018
|
|
6HTJ
 
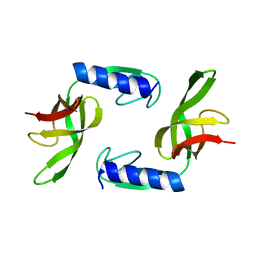 | |
2M9O
 
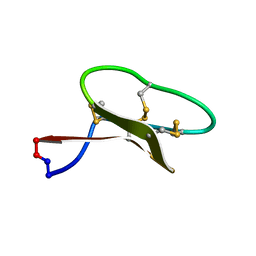 | | Solution structure of kalata B7 | | Descriptor: | Kalata-B7 | | Authors: | Daly, N, Elliott, A, Craik, D. | | Deposit date: | 2013-06-18 | | Release date: | 2013-11-20 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | Cyclotides as templates for peptide GPCR ligand design - discovery of the target receptor of the oxytocic plant peptide kalata B7
Proc.Natl.Acad.Sci.USA, 2013
|
|
