8P87
 
 | | Crystal structure of the main protease (3CLpro/Mpro) of SARS-CoV-2 obtained in presence of 5 mM X77, from an "old" crystal. | | Descriptor: | 1,2-ETHANEDIOL, 3C-like proteinase nsp5, ACETATE ION, ... | | Authors: | Costanzi, E, Demitri, N, Storici, P. | | Deposit date: | 2023-05-31 | | Release date: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Unexpected Single-Ligand Occupancy and Negative Cooperativity in the SARS-CoV-2 Main Protease.
J.Chem.Inf.Model., 64, 2024
|
|
8P54
 
 | | Crystal structure of the main protease (3CLpro/Mpro) of SARS-CoV-2 obtained in presence of 150 micromolar MG-132. | | Descriptor: | 1,2-ETHANEDIOL, 3C-like proteinase nsp5, CHLORIDE ION, ... | | Authors: | Costanzi, E, Demitri, N, Storici, P. | | Deposit date: | 2023-05-23 | | Release date: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Unexpected Single-Ligand Occupancy and Negative Cooperativity in the SARS-CoV-2 Main Protease.
J.Chem.Inf.Model., 64, 2024
|
|
8P58
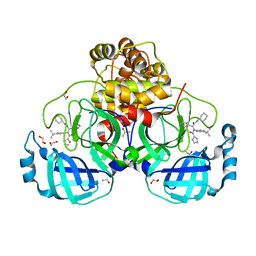 
 | | Crystal structure of the main protease (3CLpro/Mpro) of SARS-CoV-2 obtained in presence of 500 micromolar X77 enantiomer R. | | Descriptor: | 1,2-ETHANEDIOL, 3C-like proteinase nsp5, CHLORIDE ION, ... | | Authors: | Costanzi, E, Demitri, N, Storici, P. | | Deposit date: | 2023-05-23 | | Release date: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Unexpected Single-Ligand Occupancy and Negative Cooperativity in the SARS-CoV-2 Main Protease.
J.Chem.Inf.Model., 64, 2024
|
|
8P86
 
 | | Crystal structure of the main protease (3CLpro/Mpro) of SARS-CoV-2 obtained in presence of 5 mM MG-132, from an "old" crystal. | | Descriptor: | 1,2-ETHANEDIOL, 3C-like proteinase nsp5, CHLORIDE ION, ... | | Authors: | Costanzi, E, Demitri, N, Storici, P. | | Deposit date: | 2023-05-31 | | Release date: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Unexpected Single-Ligand Occupancy and Negative Cooperativity in the SARS-CoV-2 Main Protease.
J.Chem.Inf.Model., 64, 2024
|
|
4KE0
 
 | | Crystal structure of BACE1 in complex with hydroxyethylamine-macrocyclic inhibitor 13 | | Descriptor: | (3S)-3-[(1R)-2-{[(4S)-6-ethyl-3,4-dihydrospiro[chromene-2,1'-cyclobutan]-4-yl]amino}-1-hydroxyethyl]-4-azabicyclo[10.3.1]hexadeca-1(16),12,14-trien-5-one, Beta-secretase 1, GLYCEROL, ... | | Authors: | Whittington, D.A, Long, A.M, Li, V. | | Deposit date: | 2013-04-25 | | Release date: | 2013-07-03 | | Last modified: | 2013-07-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Hydroxyethylamine-based inhibitors of BACE1: P1-P3 macrocyclization can improve potency, selectivity, and cell activity.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
4K9H
 
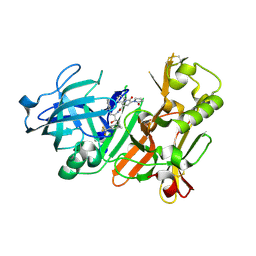 | | Bace-1 inhibitor complex | | Descriptor: | 1-cyclopentyl-N-[(2S,3R)-3-hydroxy-1-phenyl-4-{[3-(trifluoromethyl)benzyl]amino}butan-2-yl]-6-oxo-5-(2-oxopyrrolidin-1-yl)-1,6-dihydropyridine-3-carboxamide, Beta-secretase 1 | | Authors: | Jordan, S.R. | | Deposit date: | 2013-04-19 | | Release date: | 2013-07-10 | | Last modified: | 2013-07-17 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Hydroxyethylamine-based inhibitors of BACE1: P1-P3 macrocyclization can improve potency, selectivity, and cell activity.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
4K8S
 
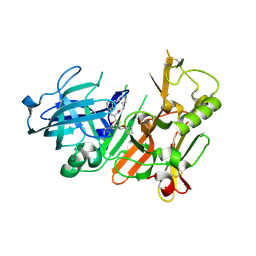 | | Hydroxyethylamine-based inhibitors of BACE1: P1-P3 macrocyclization can improve potency, selectivity, and cell activity | | Descriptor: | (3S)-3-[(1R)-2-{[(4S)-6-ethyl-3,4-dihydrospiro[chromene-2,1'-cyclobutan]-4-yl]amino}-1-hydroxyethyl]-4-azabicyclo[11.3.1]heptadeca-1(17),13,15-trien-5-one, Beta-secretase 1 | | Authors: | Jordan, S.R. | | Deposit date: | 2013-04-18 | | Release date: | 2013-07-10 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Hydroxyethylamine-based inhibitors of BACE1: P1-P3 macrocyclization can improve potency, selectivity, and cell activity.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
4KE1
 
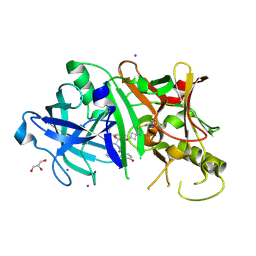 | | Crystal structure of BACE1 in complex with hydroxyethylamine-macrocyclic inhibitor 19 | | Descriptor: | (12S)-12-[(1R)-2-{[(4S)-6-ethyl-3,4-dihydrospiro[chromene-2,1'-cyclobutan]-4-yl]amino}-1-hydroxyethyl]-1,13-diazatricyclo[13.3.1.1~6,10~]icosa-6(20),7,9,15(19),16-pentaene-14,18-dione, Beta-Secretase 1, GLYCEROL, ... | | Authors: | Whittington, D.A, Long, A.M, Li, V. | | Deposit date: | 2013-04-25 | | Release date: | 2013-07-03 | | Last modified: | 2013-07-17 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Hydroxyethylamine-based inhibitors of BACE1: P1-P3 macrocyclization can improve potency, selectivity, and cell activity.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
6PGO
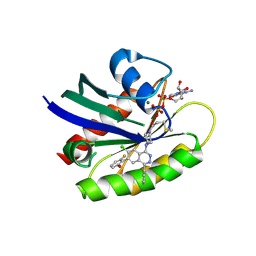 
 | | Crystal structure of human KRAS G12C covalently bound to a phthalazine inhibitor | | Descriptor: | 1-{4-[7-chloro-6-(2-fluoro-6-hydroxyphenyl)-4-phenylphthalazin-1-yl]piperazin-1-yl}propan-1-one, GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Mohr, C. | | Deposit date: | 2019-06-24 | | Release date: | 2019-12-25 | | Last modified: | 2020-01-22 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Discovery of a Covalent Inhibitor of KRASG12C(AMG 510) for the Treatment of Solid Tumors.
J.Med.Chem., 63, 2020
|
|
6PGP
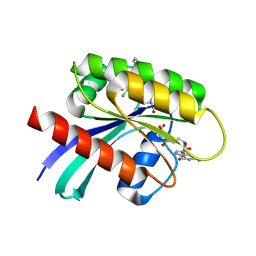 
 | |
8BPD
 
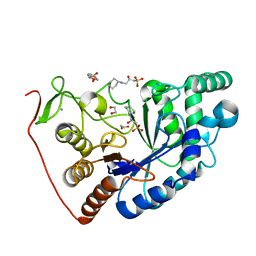 | | Structural and Functional Characterization of the Novel Endo-alpha(1,4)-Fucoidanase Mef1 from the Marine Bacterium Muricauda eckloniae | | Descriptor: | (2S)-3-(cyclohexylamino)-2-hydroxypropane-1-sulfonic acid, 1,2-ETHANEDIOL, CALCIUM ION, ... | | Authors: | Mikkelsen, M.D, Meyer, A.S, Morth, J.P. | | Deposit date: | 2022-11-16 | | Release date: | 2023-11-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and functional characterization of the novel endo-alpha (1,4)-fucoidanase Mef1 from the marine bacterium Muricauda eckloniae.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
4DH6
 
 | | Structure of Bace-1 (Beta-Secretase) in Complex with (2R)-N-((2S,3R)-1-(benzo[d][1,3]dioxol-5-yl)-3-hydroxy-4-((S)-6'-neopentyl-3',4'-dihydrospiro[cyclobutane-1,2'-pyrano[2,3-b]pyridine]-4'-ylamino)butan-2-yl)-2-methoxypropanamide | | Descriptor: | (2R)-N-[(2S,3R)-1-(1,3-benzodioxol-5-yl)-4-{[(4'S)-6'-(2,2-dimethylpropyl)-3',4'-dihydrospiro[cyclobutane-1,2'-pyrano[2,3-b]pyridin]-4'-yl]amino}-3-hydroxybutan-2-yl]-2-methoxypropanamide, Beta-secretase 1, GLYCEROL, ... | | Authors: | Sickmier, E.A. | | Deposit date: | 2012-01-27 | | Release date: | 2012-04-18 | | Last modified: | 2013-07-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Design and preparation of a potent series of hydroxyethylamine containing beta-secretase inhibitors that demonstrate robust reduction of central beta-amyloid.
J.Med.Chem., 55, 2012
|
|
7L8P
 
 | | Integrin alphaIIbbeta3 in complex with sibrafiban | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Lin, F.-Y, Springer, T.A. | | Deposit date: | 2020-12-31 | | Release date: | 2022-06-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.34995365 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
4DI2
 
 | | Crystal structure of BACE1 in complex with hydroxyethylamine inhibitor 37 | | Descriptor: | (2R)-N-{(2S,3R)-4-{[(4'S)-6'-(2,2-dimethylpropyl)-3',4'-dihydrospiro[cyclobutane-1,2'-pyrano[2,3-b]pyridin]-4'-yl]amino}-3-hydroxy-1-[3-(1,3-thiazol-2-yl)phenyl]butan-2-yl}-2-methoxypropanamide, Beta-secretase 1, GLYCEROL | | Authors: | Whittington, D.A, Long, A.M. | | Deposit date: | 2012-01-30 | | Release date: | 2012-10-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design and synthesis of potent, orally efficacious hydroxyethylamine derived beta-site amyloid precursor protein cleaving enzyme (BACE1) inhibitors.
J.Med.Chem., 55, 2012
|
|
4IO9
 
 | | Crystal structure of compound 4d bound to large ribosomal subunit (50S) from Deinococcus radiodurans | | Descriptor: | (3aS,4R,7R,8S,9S,10R,11R,13R,15R,15aR)-4-ethyl-11-methoxy-3a,7,9,11,13,15-hexamethyl-2,6,14-trioxo-10-{[3,4,6-trideoxy-3-(dimethylamino)-beta-D-xylo-hexopyranosyl]oxy}tetradecahydro-2H-oxacyclotetradecino[4,3-d][1,3]oxazol-8-yl (2R)-2-(pyridin-3-yl)pyrrolidine-1-carboxylate, 23S ribosomal RNA, 50S ribosomal protein L11, ... | | Authors: | Han, S, Marr, E.S. | | Deposit date: | 2013-01-07 | | Release date: | 2013-03-06 | | Last modified: | 2013-03-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Novel 3-O-carbamoyl erythromycin A derivatives (carbamolides) with activity against resistant staphylococcal and streptococcal isolates.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
4IOA
 
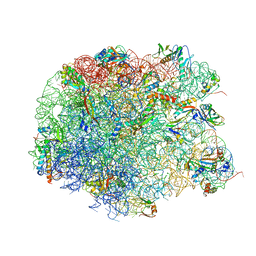 | | Crystal structure of compound 4e bound to large ribosomal subunit (50S) from Deinococcus radiodurans | | Descriptor: | (3aS,4R,7R,8S,9S,10R,11R,13R,15R,15aR)-4-ethyl-11-methoxy-3a,7,9,11,13,15-hexamethyl-2,6,14-trioxo-10-{[3,4,6-trideoxy-3-(dimethylamino)-beta-D-xylo-hexopyranosyl]oxy}tetradecahydro-2H-oxacyclotetradecino[4,3-d][1,3]oxazol-8-yl (2R)-2-[4-(acetylamino)phenyl]-2,3-dihydro-1H-pyrrole-1-carboxylate, 23S ribosomal RNA, 50S ribosomal protein L11, ... | | Authors: | Han, S, Marr, E.S. | | Deposit date: | 2013-01-07 | | Release date: | 2013-03-06 | | Last modified: | 2013-03-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Novel 3-O-carbamoyl erythromycin A derivatives (carbamolides) with activity against resistant staphylococcal and streptococcal isolates.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
4IOC
 
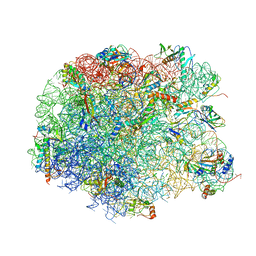 | | Crystal structure of compound 4f bound to large ribosomal subunit (50S) from Deinococcus radiodurans | | Descriptor: | (3aS,4R,7R,8S,9S,10R,11R,13R,15S,15aR)-4-ethyl-11-methoxy-3a,7,9,11,13,15-hexamethyl-2,6,14-trioxo-10-{[3,4,6-trideoxy-3-(dimethylamino)-beta-D-xylo-hexopyranosyl]oxy}tetradecahydro-2H-oxacyclotetradecino[4,3-d][1,3]oxazol-8-yl (2R)-4,4-dimethyl-2-(pyridin-3-yl)pyrrolidine-1-carboxylate, 23S ribosomal RNA, 50S ribosomal protein L11, ... | | Authors: | Han, S, Marr, E.S. | | Deposit date: | 2013-01-07 | | Release date: | 2013-03-06 | | Last modified: | 2013-03-20 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Novel 3-O-carbamoyl erythromycin A derivatives (carbamolides) with activity against resistant staphylococcal and streptococcal isolates.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
7UKO
 
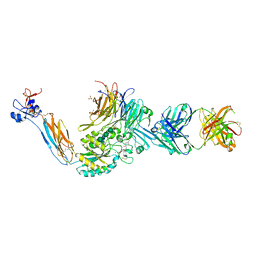 | | Integrin alaphIIBbeta3 complex with sibrafiban (Mn) | | Descriptor: | 10E5 Fab heavy chain, 10E5 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-04-01 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.604 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
7UJK
 
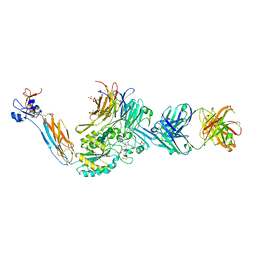 | | Integrin alaphIIBbeta3 complex with lamifiban | | Descriptor: | 10E5 Fab heavy chain, 10E5 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-03-30 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.43265581 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
7UDG
 
 | | Integrin alaphIIBbeta3 complex with lotrafiban | | Descriptor: | 10E5 Fab heavy chain, 10E5 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-03-19 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.800069 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
7UJE
 
 | | Integrin alaphIIBbeta3 complex with UR2922 in Mn2+ | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-03-30 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.499968 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
7UE0
 
 | | Integrin alaphIIBbeta3 complex with fradafiban | | Descriptor: | 10E5 Fab heavy chain, 10E5 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-03-20 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.741962 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
7UKP
 
 | | Integrin alaphIIBbeta3 complex with a gantofiban analog | | Descriptor: | 10E5 Fab heavy chain, 10E5 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-04-01 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.80112982 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
7UBR
 
 | | Integrin alaphIIBbeta3 complex with GR144053 | | Descriptor: | 10E5 Fab heavy chain, 10E5 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-03-15 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.04992747 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
7UH8
 
 | | Integrin alaphIIBbeta3 complex with roxifiban (Mn/Ca) | | Descriptor: | 10E5 Fab heavy chain, 10E5 Fab light chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Lin, F.-Y, Zhu, J, Zhu, J, Springer, T.A. | | Deposit date: | 2022-03-25 | | Release date: | 2022-08-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.750024 Å) | | Cite: | A general chemical principle for creating closure-stabilizing integrin inhibitors.
Cell, 185, 2022
|
|
