6A8P
 
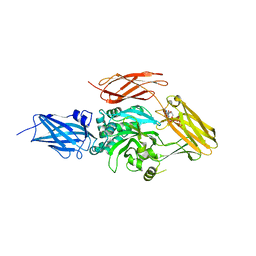 | | Transglutaminase 2 mutant G224V in complex with GTP | | Descriptor: | GUANOSINE-5'-TRIPHOSPHATE, Protein-glutamine gamma-glutamyltransferase 2 | | Authors: | Park, H.H, Ha, H.J, Kwon, S. | | Deposit date: | 2018-07-09 | | Release date: | 2018-09-26 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.537 Å) | | Cite: | Structure of natural variant transglutaminase 2 reveals molecular basis of gaining stability and higher activity.
PLoS ONE, 13, 2018
|
|
5XPC
 
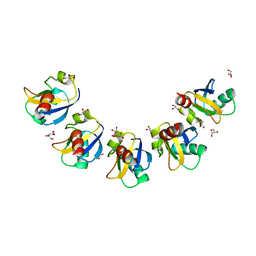 | | Crystal Structure of Drep4 CIDE domain | | Descriptor: | DNAation factor-related protein 4, GLYCEROL | | Authors: | Park, H.H, Jeong, J.H. | | Deposit date: | 2017-06-01 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | CIDE domains form functionally important higher-order assemblies for DNA fragmentation.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
7BRA
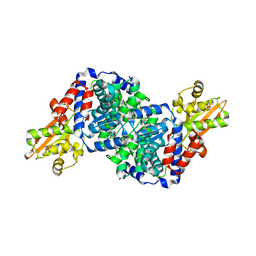 
 | | Bacillus subtilis IRG1 | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, Bacillus subtilis IRG1, SULFATE ION | | Authors: | Park, H.H, Chun, H.L. | | Deposit date: | 2020-03-27 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.785 Å) | | Cite: | Enzymatic reaction mechanism of cis-aconitate decarboxylase based on the crystal structure of IRG1 from Bacillus subtilis.
Sci Rep, 10, 2020
|
|
7BR9
 
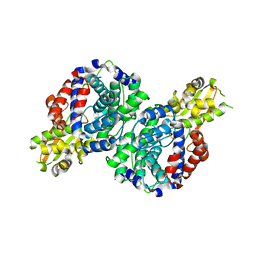 | | Crystal structure of mus musculus IRG1 | | Descriptor: | Cis-aconitate decarboxylase | | Authors: | Park, H.H, Chun, H.L. | | Deposit date: | 2020-03-27 | | Release date: | 2021-02-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | The crystal structure of mouse IRG1 suggests that cis-aconitate decarboxylase has an open and closed conformation.
Plos One, 15, 2020
|
|
7D1I
 
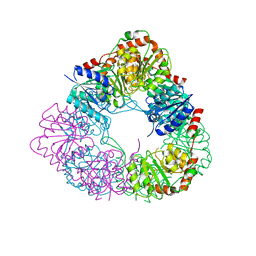 | | Crystal structure of acinetobacter baumannii MurG | | Descriptor: | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase | | Authors: | Park, H.H, Jeong, k.H. | | Deposit date: | 2020-09-14 | | Release date: | 2021-07-14 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.487 Å) | | Cite: | Putative hexameric glycosyltransferase functional unit revealed by the crystal structure of Acinetobacter baumannii MurG
Iucrj, 8, 2021
|
|
7D27
 
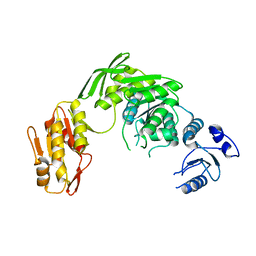 | |
7BSO
 
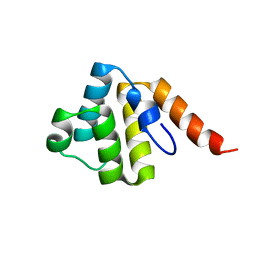 | | Crystal structure of the human NLRP9 pyrin domain | | Descriptor: | NACHT, LRR and PYD domains-containing protein 9 | | Authors: | Park, H.H, Ha, H.J. | | Deposit date: | 2020-03-31 | | Release date: | 2021-02-10 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Crystal structure of the human NLRP9 pyrin domain reveals a bent N-terminal loop that may regulate inflammasome assembly.
Febs Lett., 594, 2020
|
|
6J52
 
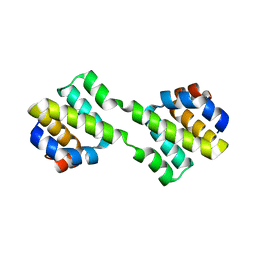 | | Crystal structure of CARD-only protein in frog virus 3 | | Descriptor: | Caspase recruitment domain-only protein | | Authors: | Park, H.H, Kwon, S. | | Deposit date: | 2019-01-10 | | Release date: | 2019-02-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.504 Å) | | Cite: | Structural transformation-mediated dimerization of caspase recruitment domain revealed by the crystal structure of CARD-only protein in frog virus 3.
J. Struct. Biol., 205, 2019
|
|
6K8H
 
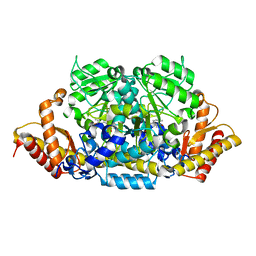 | | Crystal structure of an omega-transaminase from Sphaerobacter thermophilus | | Descriptor: | (5-HYDROXY-4,6-DIMETHYLPYRIDIN-3-YL)METHYL DIHYDROGEN PHOSPHATE, Aminotransferase class-III | | Authors: | Park, H.H, Kwon, S. | | Deposit date: | 2019-06-12 | | Release date: | 2019-10-09 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural insights into the enzyme specificity of a novel omega-transaminase from the thermophilic bacterium Sphaerobacter thermophilus.
J.Struct.Biol., 208, 2019
|
|
6KU5
 
 | | Notothenia coriiceps TRAF5 | | Descriptor: | TRAF5 | | Authors: | Park, H.H, Kim, C.M. | | Deposit date: | 2019-08-30 | | Release date: | 2020-07-08 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (3.299 Å) | | Cite: | Structural and biochemical characterization of TRAF5 from Notothenia coriiceps and its implications in fish immune cell signaling.
Fish Shellfish Immunol., 102, 2020
|
|
6KU6
 
 | | OSM1 mutant - R326A | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Fumarate reductase 2, SUCCINIC ACID | | Authors: | Park, H.H, Kim, C.M. | | Deposit date: | 2019-08-30 | | Release date: | 2020-07-08 | | Method: | X-RAY DIFFRACTION (2.007 Å) | | Cite: | Crystal Structure of the Active Site Mutant Form of Soluble Fumarate Reductase, Osm1
Crystals, 2019
|
|
6KZB
 
 | | Transglutaminase2 complexed with calcium | | Descriptor: | CALCIUM ION, GUANOSINE-5'-DIPHOSPHATE, Protein-glutamine gamma-glutamyltransferase 2 | | Authors: | Park, H.H, Kim, C.M. | | Deposit date: | 2019-09-23 | | Release date: | 2020-12-09 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.55 Å) | | Cite: | Competitive Binding of Magnesium to Calcium Binding Sites Reciprocally Regulates Transamidase and GTP Hydrolysis Activity of Transglutaminase 2.
Int J Mol Sci, 21, 2020
|
|
6IO1
 
 | |
6IZ9
 
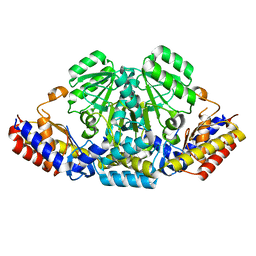 | |
4UZ0
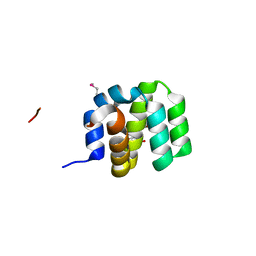 
 | | Crystal Structure of apoptosis repressor with CARD (ARC) | | Descriptor: | GLYCEROL, NUCLEOLAR PROTEIN 3 | | Authors: | Kim, S.H, Jeong, J.H, Jang, T.H, Kim, Y.G, Park, H.H. | | Deposit date: | 2014-09-04 | | Release date: | 2015-07-01 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (2.399 Å) | | Cite: | Crystal Structure of Caspase Recruiting Domain (Card) of Apoptosis Repressor with Card (Arc) and its Implication in Inhibition of Apoptosis.
Sci.Rep., 5, 2015
|
|
4D2K
 
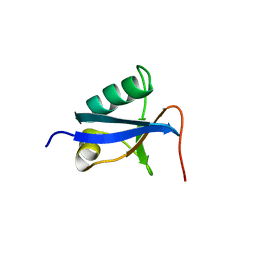 | |
7YSI
 
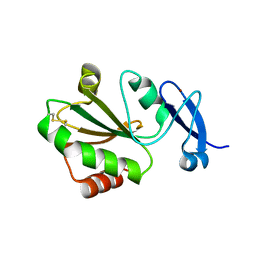 | | Crystal structure of thioredoxin 2 | | Descriptor: | Thiol disulfide reductase thioredoxin, ZINC ION | | Authors: | Chang, Y.J, Park, H.H. | | Deposit date: | 2022-08-12 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.202 Å) | | Cite: | Comparison of the structure and activity of thioredoxin 2 and thioredoxin 1 from Acinetobacter baumannii.
Iucrj, 10, 2023
|
|
4RHF
 
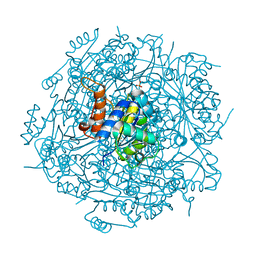 | | Crystal structure of UbiX mutant V47S from Colwellia psychrerythraea 34H | | Descriptor: | 3-octaprenyl-4-hydroxybenzoate carboxy-lyase, SULFATE ION | | Authors: | Do, H, Kim, S.J, Lee, C.W, Kim, H.-W, Park, H.H, Kim, H.M, Park, H, Park, H.J, Lee, J.H. | | Deposit date: | 2014-10-02 | | Release date: | 2015-02-18 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.764 Å) | | Cite: | Crystal structure of UbiX, an aromatic acid decarboxylase from the psychrophilic bacterium Colwellia psychrerythraea that undergoes FMN-induced conformational changes.
Sci Rep, 5, 2015
|
|
4RHE
 
 | | Crystal structure of UbiX, an aromatic acid decarboxylase from the Colwellia psychrerythraea 34H | | Descriptor: | 3-octaprenyl-4-hydroxybenzoate carboxy-lyase, FLAVIN MONONUCLEOTIDE, SULFATE ION | | Authors: | Do, H, Kim, S.J, Lee, C.W, Kim, H.-W, Park, H.H, Kim, H.M, Park, H, Park, H.J, Lee, J.H. | | Deposit date: | 2014-10-02 | | Release date: | 2015-02-18 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.003 Å) | | Cite: | Crystal structure of UbiX, an aromatic acid decarboxylase from the psychrophilic bacterium Colwellia psychrerythraea that undergoes FMN-induced conformational changes.
Sci Rep, 5, 2015
|
|
8WEQ
 
 | | p-hydroxybenzoate 3-monooxygenase | | Descriptor: | 4-hydroxybenzoate 3-monooxygenase, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Kim, S.B, Park, H.H. | | Deposit date: | 2023-09-18 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (3.29 Å) | | Cite: | Structure of p-hydroxybenzoate 3-monooxygenase
To Be Published
|
|
7ESJ
 
 | |
7EZY
 
 | | anti-CRISPR-associated Aca2 | | Descriptor: | anti-CRISPR-associated Aca2 | | Authors: | Lee, S.Y, Park, H.H. | | Deposit date: | 2021-06-02 | | Release date: | 2022-01-12 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Molecular basis of transcriptional repression ofanti-CRISPR by anti-CRISPR-associated 2
Acta Crystallogr.,Sect.D, 78, 2022
|
|
7F7P
 
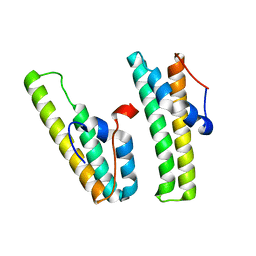 | | AcrIIC4 | | Descriptor: | anti-CRISPR protein AcrIIC4 | | Authors: | Kim, G.E, Park, H.H. | | Deposit date: | 2021-06-30 | | Release date: | 2022-05-25 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Crystal structure of the anti-CRISPR, AcrIIC4.
Protein Sci., 30, 2021
|
|
8K2Y
 
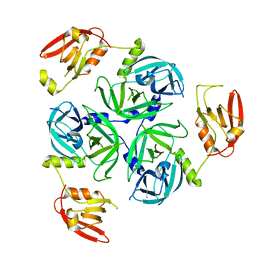 | | Crystal structure of MucD | | Descriptor: | serine endoprotease DegP-like protein MucD | | Authors: | Kim, J.H, Park, H.H. | | Deposit date: | 2023-07-14 | | Release date: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The structure of MucD from Pseudomonas syringae revealed N-terminal loop-mediated trimerization of HtrA-like serine protease.
Biochem.Biophys.Res.Commun., 688, 2023
|
|
8JQZ
 
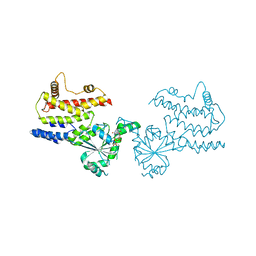 | | Crystal Structure of GppNHp-bound mIRGB10 | | Descriptor: | Immunity-related GTPase family member b10, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER | | Authors: | Ha, H.J, Park, H.H. | | Deposit date: | 2023-06-15 | | Release date: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Structural basis of IRGB10 oligomerization by GTP hydrolysis.
Front Immunol, 14, 2023
|
|
