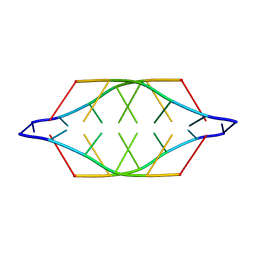
225D
 
 | |
8CT8
 
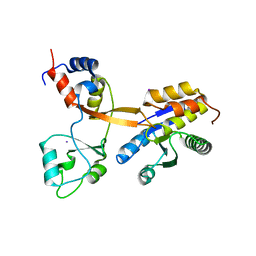 | | Crystal structure of Drosophila melanogaster PRL/CBS-pair domain complex | | 分子名称: | IODIDE ION, PRL-1 phosphatase, Unextended protein | | 著者 | Fakih, R, Goldstein, R.H, Kozlov, G, Gehring, K. | | 登録日 | 2022-05-13 | | 公開日 | 2023-03-01 | | 最終更新日 | 2024-05-22 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Burst kinetics and CNNM binding are evolutionarily conserved properties of phosphatases of regenerating liver.
J.Biol.Chem., 299, 2023
|
|
1ME0
 
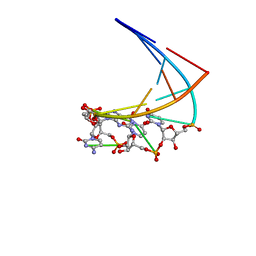 | | Chimeric hairpin with 2',5'-linked RNA loop and DNA stem | | 分子名称: | DNA/RNA (5'-D(*GP*GP*AP*C)-R(P*(U25)P*(U25)P*(C25)P*(5GP))-D(P*GP*TP*CP*C)-3') | | 著者 | Denisov, A.Y, Hannoush, R.N, Gehring, K, Damha, M.J. | | 登録日 | 2002-08-07 | | 公開日 | 2003-08-19 | | 最終更新日 | 2024-05-01 | | 実験手法 | SOLUTION NMR | | 主引用文献 | A Novel RNA Motif Based on the Structure of Unusually Stable 2',5'-Linked r(UUCG) Loops
J.AM.CHEM.SOC., 125, 2003
|
|
8F6D
 
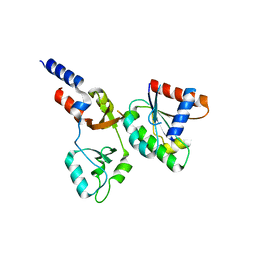 | |
2QHO
 
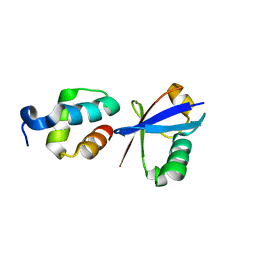 | |
8SMO
 
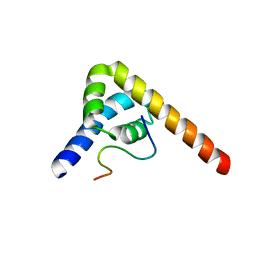 | |
6DJX
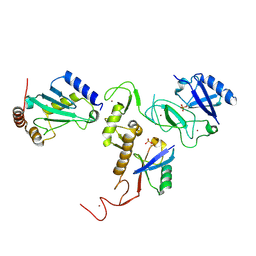 
 | | Crystal Structure of pParkin-pUb-UbcH7 complex | | 分子名称: | RBR-type E3 ubiquitin transferase,RBR-type E3 ubiquitin transferase, Ubiquitin, Ubiquitin-conjugating enzyme E2 L3, ... | | 著者 | Sauve, V, Sung, G, Trempe, J.F, Gehring, K. | | 登録日 | 2018-05-27 | | 公開日 | 2018-07-04 | | 最終更新日 | 2023-10-11 | | 実験手法 | X-RAY DIFFRACTION (4.801 Å) | | 主引用文献 | Mechanism of parkin activation by phosphorylation.
Nat. Struct. Mol. Biol., 25, 2018
|
|
6DFD
 
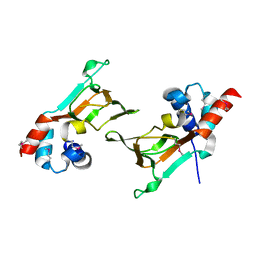 | |
7SOW
 
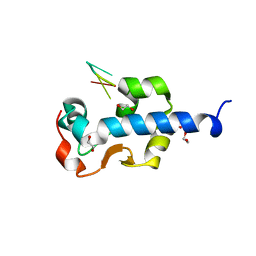 | |
7US1
 
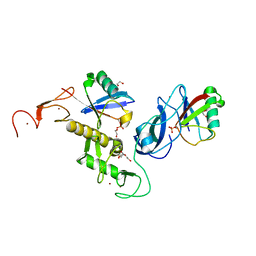 | | Structure of parkin (R0RB) bound to two phospho-ubiquitin molecules | | 分子名称: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, E3 ubiquitin-protein ligase parkin, ... | | 著者 | Fakih, R, Sauve, V, Gehring, K. | | 登録日 | 2022-04-22 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.484 Å) | | 主引用文献 | Structure of the second phosphoubiquitin-binding site in parkin.
J.Biol.Chem., 298, 2022
|
|
4F26
 
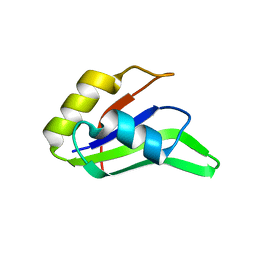 | |
4F25
 
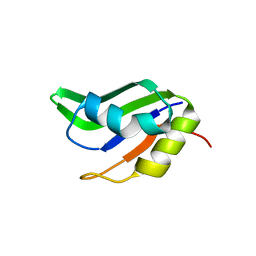 | |
6PIK
 
 | | Tetrameric cryo-EM ArnA | | 分子名称: | Bifunctional polymyxin resistance protein ArnA, UDP-4-amino-4-deoxy-L-arabinose formyltransferase | | 著者 | Yang, M, Gehring, K. | | 登録日 | 2019-06-26 | | 公開日 | 2019-07-31 | | 最終更新日 | 2024-03-20 | | 実験手法 | ELECTRON MICROSCOPY (7.8 Å) | | 主引用文献 | Cryo-electron microscopy structures of ArnA, a key enzyme for polymyxin resistance, revealed unexpected oligomerizations and domain movements.
J.Struct.Biol., 208, 2019
|
|
7SOQ
 
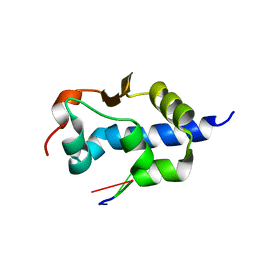 | |
7SOV
 
 | |
7SOP
 
 | |
7SOS
 
 | |
7SOR
 
 | |
7SOU
 
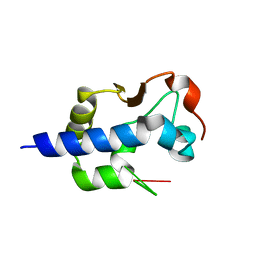 | |
7SOO
 
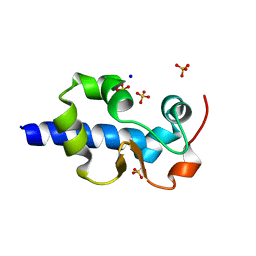 | | LaM domain of human LARP1 | | 分子名称: | Isoform 2 of La-related protein 1, SODIUM ION, SULFATE ION | | 著者 | Kozlov, G, Gehring, K. | | 登録日 | 2021-11-01 | | 公開日 | 2022-08-03 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Structural basis of 3'-end poly(A) RNA recognition by LARP1.
Nucleic Acids Res., 50, 2022
|
|
7SOT
 
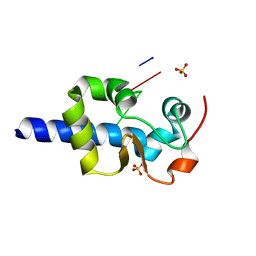 | |
6PIH
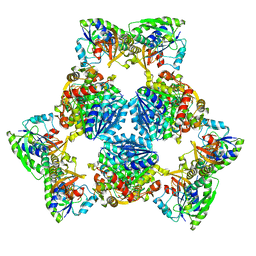 
 | | Hexameric ArnA cryo-EM structure | | 分子名称: | Bifunctional polymyxin resistance protein ArnA, UDP-4-amino-4-deoxy-L-arabinose formyltransferase | | 著者 | Yang, M, Gehring, K. | | 登録日 | 2019-06-26 | | 公開日 | 2019-07-31 | | 最終更新日 | 2024-03-13 | | 実験手法 | ELECTRON MICROSCOPY (6.6 Å) | | 主引用文献 | Cryo-electron microscopy structures of ArnA, a key enzyme for polymyxin resistance, revealed unexpected oligomerizations and domain movements.
J.Struct.Biol., 208, 2019
|
|
5TDA
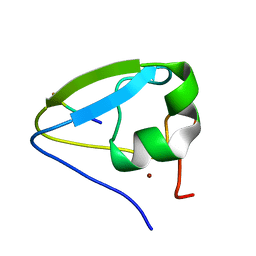 
 | |
8GK6
 
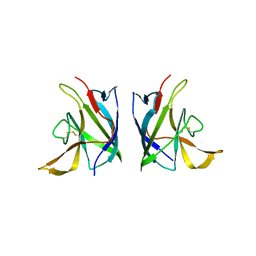 | |
5BU1
 
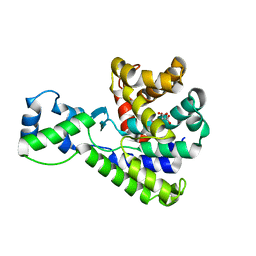 | | Structure of the truncated C-terminal domain of lpg1496 from Legionella pneumophila | | 分子名称: | LPG1496, MALONATE ION | | 著者 | Wong, K, Kozlov, G, Gehring, K, Montreal-Kingston Bacterial Structural Genomics Initiative (BSGI) | | 登録日 | 2015-06-03 | | 公開日 | 2015-08-26 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Structure of the Legionella Effector, lpg1496, Suggests a Role in Nucleotide Metabolism.
J.Biol.Chem., 290, 2015
|
|
