6U9R
 
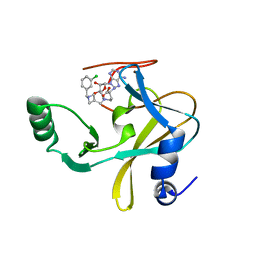 | | MLL1 SET N3861I/Q3867L bound to inhibitor 12 (TC-5140) | | 分子名称: | 5'-{[(3S)-3-amino-3-carboxypropyl]({1-[(3-chlorophenyl)methyl]azetidin-3-yl}methyl)amino}-5'-deoxyadenosine, Histone-lysine N-methyltransferase, ZINC ION | | 著者 | Petrunak, E.M, Stuckey, J.A. | | 登録日 | 2019-09-09 | | 公開日 | 2020-07-01 | | 最終更新日 | 2023-10-11 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Discovery of Potent Small-Molecule Inhibitors of MLL Methyltransferase.
Acs Med.Chem.Lett., 11, 2020
|
|
5XZR
 
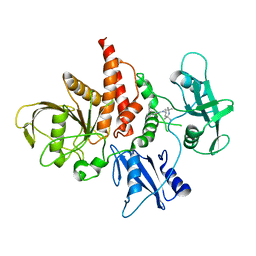 | | The atomic structure of SHP2 E76A mutant in complex with allosteric inhibitor 9b | | 分子名称: | 4-(3-phenylphenyl)-N-(2,2,6,6-tetramethylpiperidin-4-yl)-1,3-thiazol-2-amine, Tyrosine-protein phosphatase non-receptor type 11 | | 著者 | Li, D, Xie, J, Zhu, J, Liu, C. | | 登録日 | 2017-07-13 | | 公開日 | 2017-12-13 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Allosteric Inhibitors of SHP2 with Therapeutic Potential for Cancer Treatment.
J. Med. Chem., 60, 2017
|
|
5YKG
 
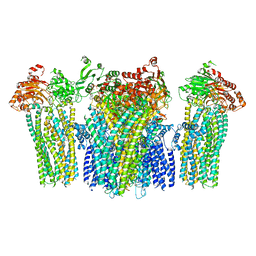 | |
5YDP
 
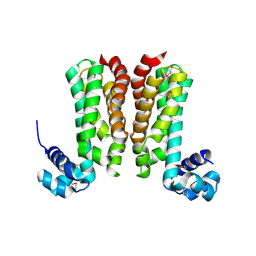 | |
8JLX
 
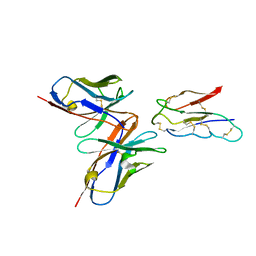 | | CCHFV envelope protein Gc in complex with Gc13 | | 分子名称: | Glycoprotein C,CCHFV Gc fusion loops, Mouse antibody Gc13 heavy chain, Mouse antibody Gc13 light chain | | 著者 | Chong, T, Cao, S. | | 登録日 | 2023-06-03 | | 公開日 | 2024-01-24 | | 最終更新日 | 2024-11-20 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | Neutralizing monoclonal antibodies against the Gc fusion loop region of Crimean-Congo hemorrhagic fever virus.
Plos Pathog., 20, 2024
|
|
5YBI
 
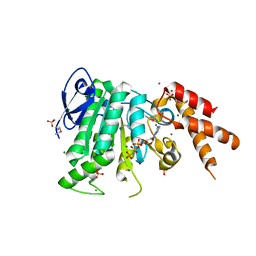 | | Structure of the bacterial pathogens ATPase with substrate AMPPNP | | 分子名称: | MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, Probable ATP synthase SpaL/MxiB, ... | | 著者 | Mu, Z.X, Gao, X.P, Cui, S. | | 登録日 | 2017-09-05 | | 公開日 | 2018-06-20 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (2.268 Å) | | 主引用文献 | Structural Insight Into Conformational Changes Induced by ATP Binding in a Type III Secretion-Associated ATPase FromShigella flexneri.
Front Microbiol, 9, 2018
|
|
5YKE
 
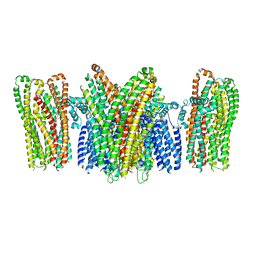 | |
6U9M
 
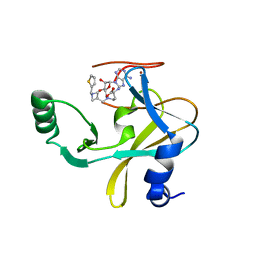 | | MLL1 SET N3861I/Q3867L bound to inhibitor 16 (TC-5109) | | 分子名称: | 5'-{[(3S)-3-amino-3-carboxypropyl]({1-[(thiophen-2-yl)methyl]azetidin-3-yl}methyl)amino}-5'-deoxyadenosine, GLYCEROL, Histone-lysine N-methyltransferase, ... | | 著者 | Petrunak, E.M, Stuckey, J.A. | | 登録日 | 2019-09-09 | | 公開日 | 2020-07-01 | | 最終更新日 | 2023-10-11 | | 実験手法 | X-RAY DIFFRACTION (2.05 Å) | | 主引用文献 | Discovery of Potent Small-Molecule Inhibitors of MLL Methyltransferase.
Acs Med.Chem.Lett., 11, 2020
|
|
5YTX
 
 | |
8JSL
 
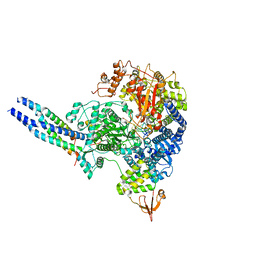 | | The structure of EBOV L-VP35-RNA complex | | 分子名称: | Polymerase cofactor VP35, RNA-directed RNA polymerase L, The leader sequence of EBOV, ... | | 著者 | Qi, P, Yi, S. | | 登録日 | 2023-06-20 | | 公開日 | 2023-09-27 | | 最終更新日 | 2023-11-01 | | 実験手法 | ELECTRON MICROSCOPY (2.95 Å) | | 主引用文献 | Molecular mechanism of de novo replication by the Ebola virus polymerase.
Nature, 622, 2023
|
|
8JSM
 
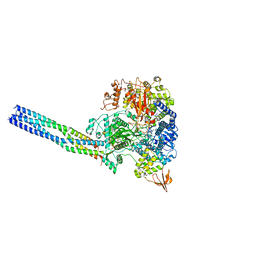 | | The structure of EBOV L-VP35-RNA complex (conformation 1) | | 分子名称: | Polymerase cofactor VP35, RNA-directed RNA polymerase L, The leader sequence of EBOV genome., ... | | 著者 | Qi, P, Yi, S. | | 登録日 | 2023-06-20 | | 公開日 | 2023-09-27 | | 最終更新日 | 2023-11-01 | | 実験手法 | ELECTRON MICROSCOPY (3.3 Å) | | 主引用文献 | Molecular mechanism of de novo replication by the Ebola virus polymerase.
Nature, 622, 2023
|
|
8JSN
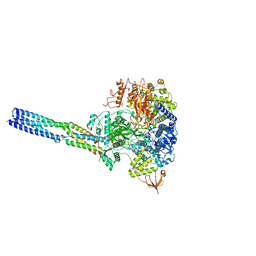 
 | | The structure of EBOV L-VP35-RNA complex (conformation 2) | | 分子名称: | Polymerase cofactor VP35, RNA-directed RNA polymerase L, The leader sequence of EBOV genome, ... | | 著者 | Qi, P, Yi, S. | | 登録日 | 2023-06-20 | | 公開日 | 2023-09-27 | | 最終更新日 | 2023-11-01 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | Molecular mechanism of de novo replication by the Ebola virus polymerase.
Nature, 622, 2023
|
|
5YTT
 
 | |
5YTV
 
 | |
5YTS
 | |
5YW9
 
 | |
5YWC
 
 | |
5YWB
 
 | |
5YW8
 
 | |
5ZT1
 
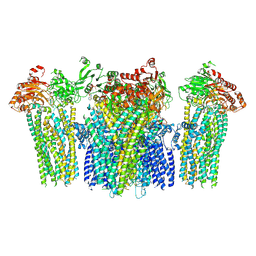
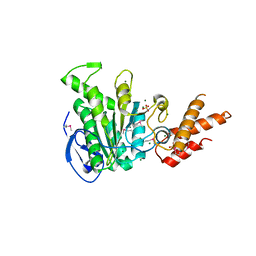 | | Structure of the bacterial pathogens ATPase with substrate ATP gamma S | | 分子名称: | MAGNESIUM ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Probable ATP synthase SpaL/MxiB, ... | | 著者 | Gao, X.P, Mu, Z.X, Cui, S. | | 登録日 | 2018-05-01 | | 公開日 | 2018-05-16 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (3.114 Å) | | 主引用文献 | Structural Insight Into Conformational Changes Induced by ATP Binding in a Type III Secretion-Associated ATPase FromShigella flexneri
Front Microbiol, 9, 2018
|
|
8JLW
 
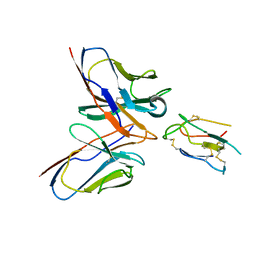 | | CCHFV envelope protein Gc in complex with Gc8 | | 分子名称: | Glycoprotein C,CCHFV envelope protein Gc fusion loops, Mouse antibody Gc8 heavy chain, Mouse antibody Gc8 light chain | | 著者 | Chong, T, Cao, S. | | 登録日 | 2023-06-03 | | 公開日 | 2024-01-24 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | Neutralizing monoclonal antibodies against the Gc fusion loop region of Crimean-Congo hemorrhagic fever virus.
Plos Pathog., 20, 2024
|
|
6WAA
 
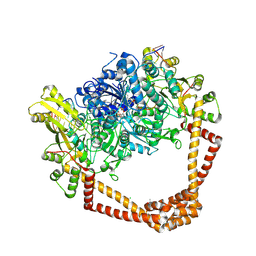 | | K. pneumoniae Topoisomerase IV (ParE-ParC) in complex with DNA and compound 34 (7-[(1S,5R)-1-amino-3-azabicyclo[3.1.0]hexan-3-yl]-4-(aminomethyl)-1-cyclopropyl-3,6-difluoro-8-methylquinolin-2(1H)-one) | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, 7-[(1S,5R)-1-amino-3-azabicyclo[3.1.0]hexan-3-yl]-4-(aminomethyl)-1-cyclopropyl-3,6-difluoro-8-methylquinolin-2(1H)-one, CHLORIDE ION, ... | | 著者 | Noeske, J, Shu, W, Bellamacina, C. | | 登録日 | 2020-03-24 | | 公開日 | 2020-07-15 | | 最終更新日 | 2024-12-25 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | Topoisomerase Inhibitors Addressing Fluoroquinolone Resistance in Gram-Negative Bacteria.
J.Med.Chem., 63, 2020
|
|
8J6T
 
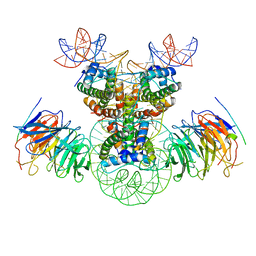 | | Cryo-EM structure of the double CAF-1 bound right-handed Di-tetrasome | | 分子名称: | Chromatin assembly factor 1 subunit A, Chromatin assembly factor 1 subunit B, Histone H3.1, ... | | 著者 | Liu, C.P, Yu, Z.Y, Xu, R.M. | | 登録日 | 2023-04-26 | | 公開日 | 2023-08-16 | | 最終更新日 | 2023-09-06 | | 実験手法 | ELECTRON MICROSCOPY (6.6 Å) | | 主引用文献 | Structural insights into histone binding and nucleosome assembly by chromatin assembly factor-1.
Science, 381, 2023
|
|
8J6S
 
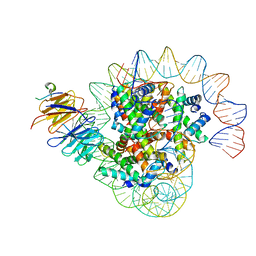 | | Cryo-EM structure of the single CAF-1 bound right-handed Di-tetrasome | | 分子名称: | Chromatin assembly factor 1 subunit A, Chromatin assembly factor 1 subunit B, Histone H3.1, ... | | 著者 | Liu, C.P, Yu, Z.Y, Xu, R.M. | | 登録日 | 2023-04-26 | | 公開日 | 2023-08-16 | | 最終更新日 | 2023-09-06 | | 実験手法 | ELECTRON MICROSCOPY (3.8 Å) | | 主引用文献 | Structural insights into histone binding and nucleosome assembly by chromatin assembly factor-1.
Science, 381, 2023
|
|
8IQF
 
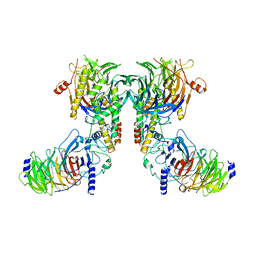 | | Cryo-EM structure of the dimeric human CAF1-H3-H4 complex | | 分子名称: | Chromatin assembly factor 1 subunit A, Chromatin assembly factor 1 subunit B, Histone H3.1, ... | | 著者 | Liu, C.P, Yu, Z.Y, Xu, R.M. | | 登録日 | 2023-03-16 | | 公開日 | 2023-08-16 | | 最終更新日 | 2023-09-06 | | 実験手法 | ELECTRON MICROSCOPY (4.6 Å) | | 主引用文献 | Structural insights into histone binding and nucleosome assembly by chromatin assembly factor-1.
Science, 381, 2023
|
|
