6APR
 | |
5ZNY
 
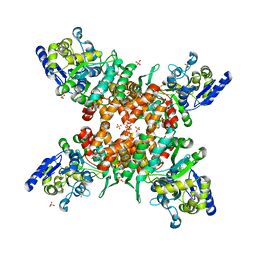 | | Structure of mDR3_DD-C363G with MBP tag | | 分子名称: | Maltose-binding periplasmic protein,Tumor necrosis factor receptor superfamily, member 25, SULFATE ION | | 著者 | Yin, X, Jin, T. | | 登録日 | 2018-04-11 | | 公開日 | 2019-04-17 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.74 Å) | | 主引用文献 | Crystal structure and activation mechanism of DR3 death domain.
Febs J., 286, 2019
|
|
5ZNZ
 
 | | Structure of mDR3 DD with MBP tag mutant-I387V | | 分子名称: | Maltose-binding periplasmic protein,Tumor necrosis factor receptor superfamily, member 25, SULFATE ION | | 著者 | Jin, T, Yin, X. | | 登録日 | 2018-04-12 | | 公開日 | 2019-04-17 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.55 Å) | | 主引用文献 | Crystal structure and activation mechanism of DR3 death domain.
Febs J., 286, 2019
|
|
5YEV
 
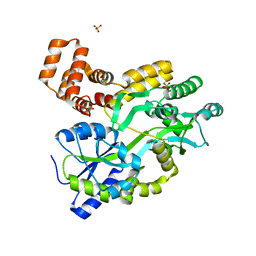 | | Murine DR3 death domain | | 分子名称: | SULFATE ION, TNFRSF25 death domain | | 著者 | Yin, X, Jin, T. | | 登録日 | 2017-09-19 | | 公開日 | 2018-09-19 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Crystal structure and activation mechanism of DR3 death domain.
Febs J., 286, 2019
|
|
2BKJ
 
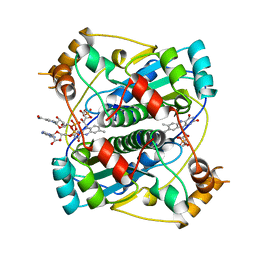 | |
2ER9
 
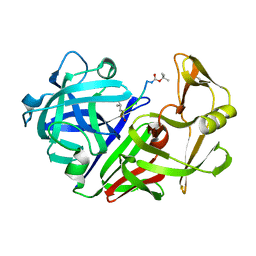 | | X-RAY STUDIES OF ASPARTIC PROTEINASE-STATINE INHIBITOR COMPLEXES. | | 分子名称: | ENDOTHIAPEPSIN, L363,564 | | 著者 | Cooper, J.B, Foundling, S.I, Boger, J, Blundell, T.L. | | 登録日 | 1990-10-20 | | 公開日 | 1991-01-15 | | 最終更新日 | 2017-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | X-ray studies of aspartic proteinase-statine inhibitor complexes.
Biochemistry, 28, 1989
|
|
2ER0
 
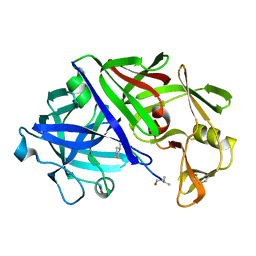 | | X-RAY STUDIES OF ASPARTIC PROTEINASE-STATINE INHIBITOR COMPLEXES | | 分子名称: | ENDOTHIAPEPSIN, L364,099 | | 著者 | Cooper, J.B, Foundling, S.I, Boger, J, Blundell, T.L. | | 登録日 | 1990-10-20 | | 公開日 | 1991-01-15 | | 最終更新日 | 2017-11-29 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | X-ray studies of aspartic proteinase-statine inhibitor complexes.
Biochemistry, 28, 1989
|
|
7C2X
 
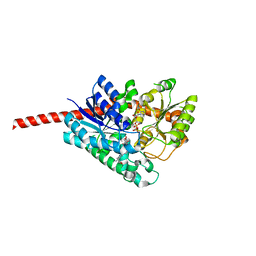 | | Crystal Structure of Glycyrrhiza uralensis UGT73P12 complexed with glycyrrhetinic acid 3-O-monoglucuronide | | 分子名称: | (2~{S},3~{S},4~{S},5~{R},6~{R})-6-[[(3~{S},4~{a}~{R},6~{a}~{R},6~{b}~{S},8~{a}~{S},11~{S},12~{a}~{R},14~{a}~{R},14~{b}~{S})-11-carboxy-4,4,6~{a},6~{b},8~{a},11,14~{b}-heptamethyl-14-oxidanylidene-2,3,4~{a},5,6,7,8,9,10,12,12~{a},14~{a}-dodecahydro-1~{H}-picen-3-yl]oxy]-3,4,5-tris(oxidanyl)oxane-2-carboxylic acid, Glycosyltransferase, URIDINE-5'-DIPHOSPHATE | | 著者 | Ren, J. | | 登録日 | 2020-05-09 | | 公開日 | 2021-05-12 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.89 Å) | | 主引用文献 | Crystal Structure of Glycyrrhiza uralensis UGT73P12 complexed with glycyrrhetinic acid 3-O-monoglucuronide
To Be Published
|
|
7DVM
 
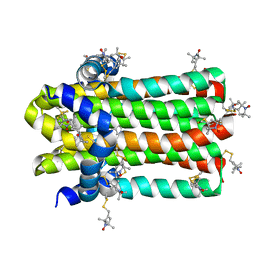 | |
4APE
 
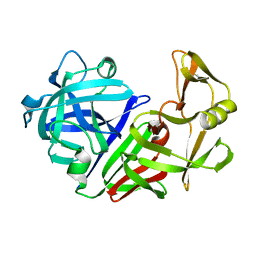 | | THE ACTIVE SITE OF ASPARTIC PROTEINASES | | 分子名称: | ENDOTHIAPEPSIN | | 著者 | Pearl, L.H, Sewell, B.T, Jenkins, J.A, Cooper, J.B, Blundell, T.L. | | 登録日 | 1986-06-09 | | 公開日 | 1986-07-14 | | 最終更新日 | 2019-10-16 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | The Active Site of Aspartic Proteinases
FEBS Lett., 174, 1984
|
|
6JIQ
 
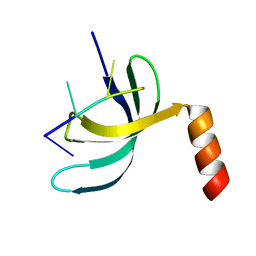 | | Crystal structure of Streptococcus pneumoniae SP_0782 (residues 7-79) in complex with single-stranded DNA dT6 | | 分子名称: | DNA (5'-D(*TP*TP*TP*TP*T)-3'), SP_0782 | | 著者 | Fang, X, Lu, G, Li, S, Zhu, J, Yang, Y, Gong, P. | | 登録日 | 2019-02-22 | | 公開日 | 2019-11-27 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.67 Å) | | 主引用文献 | Structural insight into the length-dependent binding of ssDNA by SP_0782 from Streptococcus pneumoniae, reveals a divergence in the DNA-binding interface of PC4-like proteins.
Nucleic Acids Res., 48, 2020
|
|
6JIP
 
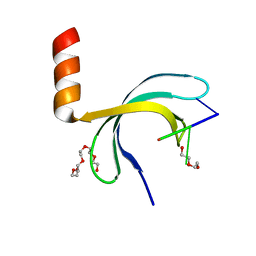 | | Crystal structure of Streptococcus pneumoniae SP_0782 (residues 7-79) in complex with single-stranded DNA dT6 | | 分子名称: | DNA (5'-D(*TP*TP*TP*TP*T)-3'), PENTAETHYLENE GLYCOL, SP_0782 | | 著者 | Fang, X, Lu, G, Li, S, Zhu, J, Yang, Y, Gong, P. | | 登録日 | 2019-02-22 | | 公開日 | 2019-11-27 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (1.659 Å) | | 主引用文献 | Structural insight into the length-dependent binding of ssDNA by SP_0782 from Streptococcus pneumoniae, reveals a divergence in the DNA-binding interface of PC4-like proteins.
Nucleic Acids Res., 48, 2020
|
|
6IYN
 
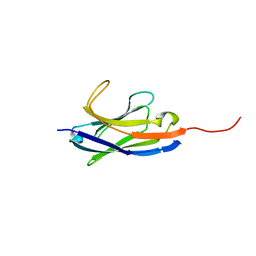 | | Solution structure of camelid nanobody Nb26 against aflatoxin B1 | | 分子名称: | Nb26 | | 著者 | Nie, Y, He, T, Zhu, J, Li, S.L, Hu, R, Yang, Y.H. | | 登録日 | 2018-12-17 | | 公開日 | 2019-01-23 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Chemical shift assignments of a camelid nanobody against aflatoxin B1.
Biomol NMR Assign, 13, 2019
|
|
6IGZ
 | | Structure of PSI-LHCI | | 分子名称: | (3S,5R,6S,3'S,5'R,6'S)-5,6,5',6'-DIEPOXY-5,6,5',6'- TETRAHYDRO-BETA,BETA-CAROTENE-3,3'-DIOL, (6'R,11cis,11'cis,13cis,15cis)-4',5'-didehydro-5',6'-dihydro-beta,beta-carotene, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, ... | | 著者 | Xiong, P, Xiaochun, Q. | | 登録日 | 2018-09-27 | | 公開日 | 2019-02-13 | | 最終更新日 | 2020-07-29 | | 実験手法 | ELECTRON MICROSCOPY (3.49 Å) | | 主引用文献 | Structure of a green algal photosystem I in complex with a large number of light-harvesting complex I subunits.
Nat Plants, 5, 2019
|
|
6K3H
 
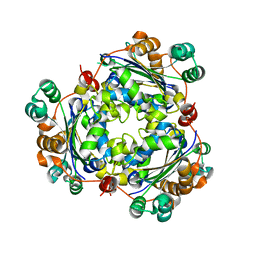 | |
6LMR
 
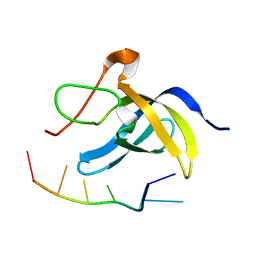 | |
6LMS
 
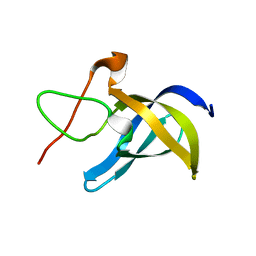 | |
6LRM
 
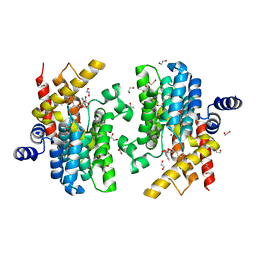 | | Crystal structure of PDE4D catalytic domain in complex with arctigenin | | 分子名称: | 1,2-ETHANEDIOL, Arctigenin, MAGNESIUM ION, ... | | 著者 | Zhang, X.L, Li, M.J, Xu, Y.C. | | 登録日 | 2020-01-16 | | 公開日 | 2021-04-14 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.45 Å) | | 主引用文献 | Identification of phosphodiesterase-4 as the therapeutic target of arctigenin in alleviating psoriatic skin inflammation.
J Adv Res, 33, 2021
|
|
6LK8
 
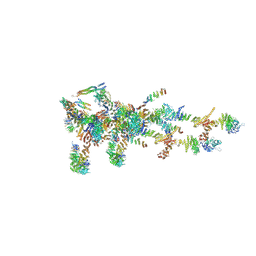 | | Structure of Xenopus laevis Cytoplasmic Ring subunit. | | 分子名称: | GATOR complex protein SEC13, MGC154553 protein, MGC83295 protein, ... | | 著者 | Shi, Y, Huang, G, Yan, C, Zhang, Y. | | 登録日 | 2019-12-18 | | 公開日 | 2021-07-21 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (5.5 Å) | | 主引用文献 | Structure of the cytoplasmic ring of the Xenopus laevis nuclear pore complex by cryo-electron microscopy single particle analysis.
Cell Res., 30, 2020
|
|
5ER2
 
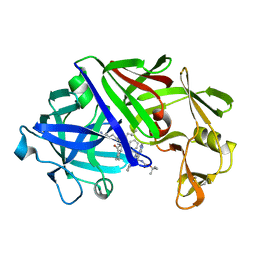 | | High-resolution X-ray diffraction study of the complex between endothiapepsin and an oligopeptide inhibitor. the analysis of the inhibitor binding and description of the rigid body shift in the enzyme | | 分子名称: | 6-ammonio-N-{[(2R,3R)-3-{[N-(tert-butoxycarbonyl)-L-phenylalanyl-3-(1H-imidazol-3-ium-4-yl)-L-alanyl]amino}-4-cyclohexyl-2-hydroxybutyl](2-methylpropyl)carbamoyl}-L-norleucyl-L-phenylalanine, ENDOTHIAPEPSIN | | 著者 | Sali, A, Veerapandian, B, Cooper, J.B, Foundling, S.I, Hoover, D.J, Blundell, T.L. | | 登録日 | 1991-01-02 | | 公開日 | 1991-04-15 | | 最終更新日 | 2017-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | High-resolution X-ray diffraction study of the complex between endothiapepsin and an oligopeptide inhibitor: the analysis of the inhibitor binding and description of the rigid body shift in the enzyme.
EMBO J., 8, 1989
|
|
5ER1
 
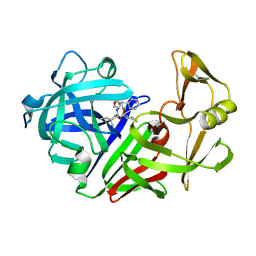 | |
7W72
 
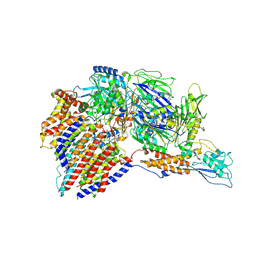 | | Structure of a human glycosylphosphatidylinositol (GPI) transamidase | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, GPI transamidase component PIG-S, ... | | 著者 | Zhang, H, Su, J, Li, B, Gao, Y, Zhang, X.C, Zhao, Y. | | 登録日 | 2021-12-02 | | 公開日 | 2022-02-16 | | 最終更新日 | 2022-03-30 | | 実験手法 | ELECTRON MICROSCOPY (3.1 Å) | | 主引用文献 | Structure of human glycosylphosphatidylinositol transamidase.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7WB4
 
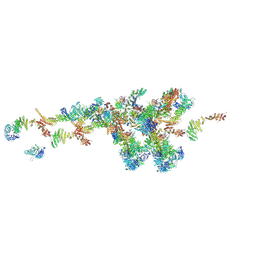 | | Cryo-EM structure of the NR subunit from X. laevis NPC | | 分子名称: | GATOR complex protein SEC13, MGC154553 protein, MGC83295 protein, ... | | 著者 | Huang, G, Zhan, X, Shi, Y. | | 登録日 | 2021-12-15 | | 公開日 | 2022-03-02 | | 最終更新日 | 2024-06-26 | | 実験手法 | ELECTRON MICROSCOPY (5.6 Å) | | 主引用文献 | Cryo-EM structure of the nuclear ring from Xenopus laevis nuclear pore complex.
Cell Res., 32, 2022
|
|
7W9N
 
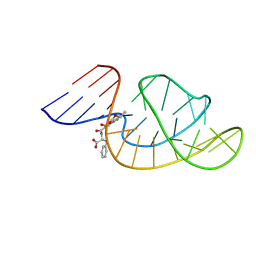 | | THE STRUCTURE OF OBA33-OTA COMPLEX | | 分子名称: | (2~{S})-2-[[(3~{R})-5-chloranyl-3-methyl-8-oxidanyl-1-oxidanylidene-3,4-dihydroisochromen-7-yl]carbonylamino]-3-phenyl-propanoic acid, OTA DNA APTAMER (33-MER) | | 著者 | Xu, G.H, Li, C.G. | | 登録日 | 2021-12-10 | | 公開日 | 2022-01-19 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Structural Insights into the Mechanism of High-Affinity Binding of Ochratoxin A by a DNA Aptamer.
J.Am.Chem.Soc., 144, 2022
|
|
7X9E
 
 | | Crystal structure of the 76E1 Fab in complex with a SARS-CoV-2 spike peptide | | 分子名称: | 76E1 Fab Heavy Chain, 76E1 Fab Light Chain, Spike peptide | | 著者 | Chen, X, Zhang, T, Ding, J, Sun, X, Sun, B. | | 登録日 | 2022-03-15 | | 公開日 | 2022-05-11 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Neutralization mechanism of a human antibody with pan-coronavirus reactivity including SARS-CoV-2.
Nat Microbiol, 7, 2022
|
|
