7E8D
 
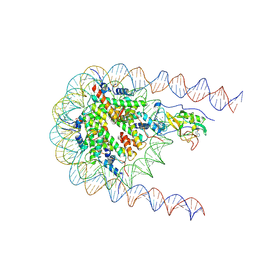 | | NSD2 E1099K mutant bound to nucleosome | | 分子名称: | DNA (185-MER), Histone H2A type 1, Histone H2B type 1-J, ... | | 著者 | Sengoku, T, Sato, K, Nishizawa, T, Nureki, O, Ogata, K. | | 登録日 | 2021-03-01 | | 公開日 | 2021-11-10 | | 最終更新日 | 2025-06-18 | | 実験手法 | ELECTRON MICROSCOPY (2.8 Å) | | 主引用文献 | Structural basis of the regulation of the normal and oncogenic methylation of nucleosomal histone H3 Lys36 by NSD2.
Nat Commun, 12, 2021
|
|
5J5D
 
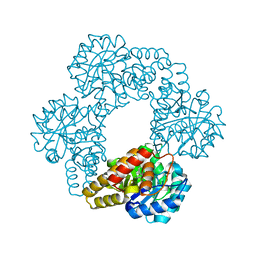 | |
7BWE
 
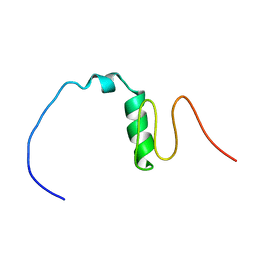 | |
7BWO
 | |
6W1J
 
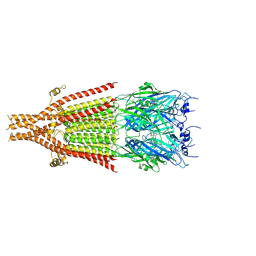 | | Cryo-EM structure of 5HT3A receptor in presence of Alosetron | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(4-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-hydroxytryptamine receptor 3A, ... | | 著者 | Basak, S, Chakrapani, S. | | 登録日 | 2020-03-04 | | 公開日 | 2021-01-13 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (2.92 Å) | | 主引用文献 | High-resolution structures of multiple 5-HT 3A R-setron complexes reveal a novel mechanism of competitive inhibition.
Elife, 9, 2020
|
|
6W1M
 
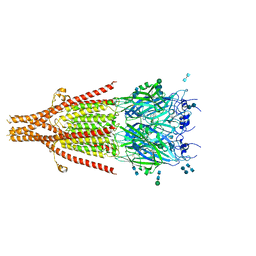
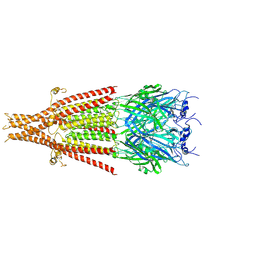 | | Cryo-EM structure of 5HT3A receptor in presence of Ondansetron | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(4-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-hydroxytryptamine receptor 3A, ... | | 著者 | Basak, S, Chakrapani, S. | | 登録日 | 2020-03-04 | | 公開日 | 2021-01-13 | | 最終更新日 | 2024-11-20 | | 実験手法 | ELECTRON MICROSCOPY (3.06 Å) | | 主引用文献 | High-resolution structures of multiple 5-HT 3A R-setron complexes reveal a novel mechanism of competitive inhibition.
Elife, 9, 2020
|
|
6W1Y
 
 | | Cryo-EM structure of 5HT3A receptor in presence of Palonosetron | | 分子名称: | (3~{a}~{S})-2-[(3~{S})-1-azabicyclo[2.2.2]octan-3-yl]-3~{a},4,5,6-tetrahydro-3~{H}-benzo[de]isoquinolin-1-one, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(4-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Basak, S, Chakrapani, S. | | 登録日 | 2020-03-04 | | 公開日 | 2021-01-13 | | 最終更新日 | 2024-11-06 | | 実験手法 | ELECTRON MICROSCOPY (3.35 Å) | | 主引用文献 | High-resolution structures of multiple 5-HT 3A R-setron complexes reveal a novel mechanism of competitive inhibition.
Elife, 9, 2020
|
|
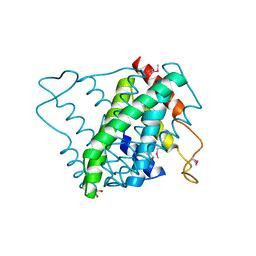
3NL9
 
 | |
3HSA
 
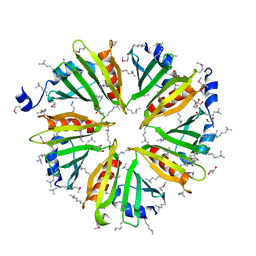 | |
3GF8
 
 | |
4BQP
 
 | | Mtb InhA complex with Methyl-thiazole compound 7 | | 分子名称: | (1S)-1-(5-{[1-(2,6-DIFLUOROBENZYL)-1H-PYRAZOL-3-YL]AMINO}-1,3,4-THIADIAZOL-2-YL)-1-(4-METHYL-1,3-THIAZOL-2-YL)ETHANOL, ENOYL-[ACYL-CARRIER-PROTEIN] REDUCTASE [NADH], NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | 著者 | Read, J.A, Gingell, H, Madhavapeddi, P, Shirude, P.S. | | 登録日 | 2013-05-31 | | 公開日 | 2013-12-11 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (1.89 Å) | | 主引用文献 | Methyl-Thiazoles: A Novel Mode of Inhibition with the Potential to Develop Novel Inhibitors Targeting Inha in Mycobacterium Tuberculosis.
J.Med.Chem., 56, 2013
|
|
3H0N
 
 | |
3H41
 
 | |
4BQR
 
 | | Mtb InhA complex with Methyl-thiazole compound 11 | | 分子名称: | (NZ)-2-[2,6-bis(fluoranyl)phenyl]-N-[5-[(1S)-1-(4-methyl-1,3-thiazol-2-yl)-1-oxidanyl-ethyl]-3H-1,3,4-thiadiazol-2-ylidene]ethanamide, ENOYL-[ACYL-CARRIER-PROTEIN] REDUCTASE [NADH], NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | 著者 | Read, J.A, Gingell, H, Madhavapeddi, P, Shirude, P.S. | | 登録日 | 2013-05-31 | | 公開日 | 2013-12-11 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (2.05 Å) | | 主引用文献 | Methyl-Thiazoles: A Novel Mode of Inhibition with the Potential to Develop Novel Inhibitors Targeting Inha in Mycobacterium Tuberculosis.
J.Med.Chem., 56, 2013
|
|
4D4O
 
 | |
4D4P
 
 | |
3L5O
 
 | |
8OUJ
 
 | |
8OUD
 
 | |
8OUH
 
 | | Complex of human ASCT2 with Syncytin-1 | | 分子名称: | ALANINE, Neutral amino acid transporter B(0), Syncytin-1 | | 著者 | Khare, S, Reyes, N. | | 登録日 | 2023-04-23 | | 公開日 | 2024-05-01 | | 最終更新日 | 2024-11-06 | | 実験手法 | ELECTRON MICROSCOPY (2.62 Å) | | 主引用文献 | Receptor-recognition and antiviral mechanisms of retrovirus-derived human proteins.
Nat.Struct.Mol.Biol., 31, 2024
|
|
3HBZ
 
 | |
5VF5
 
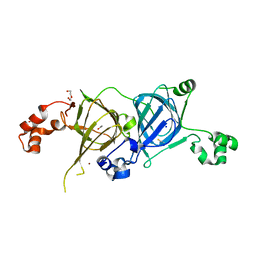 | | Crystal structure of the vicilin from Solanum melongena, re-refinement | | 分子名称: | ACETATE ION, COPPER (II) ION, DI(HYDROXYETHYL)ETHER, ... | | 著者 | Porebski, P.J, Wlodawer, A, Dauter, Z, Minor, W, Stanfield, R, Jaskolski, M, Pozharski, E, Weichenberger, C.X, Rupp, B. | | 登録日 | 2017-04-06 | | 公開日 | 2017-12-06 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.49 Å) | | 主引用文献 | Detect, correct, retract: How to manage incorrect structural models.
FEBS J., 285, 2018
|
|
4BFH
 
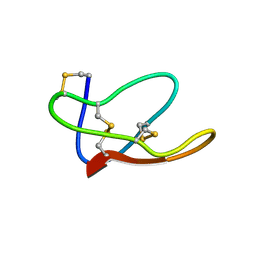 | |
4QBK
 
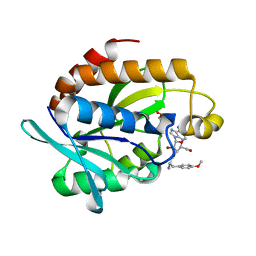 | | Crystal structure of the complex of Peptidyl-tRNA hydrolase from Pseudomonas aeruginosa with amino acyl-tRNA analogue at 1.77 Angstrom resolution | | 分子名称: | 3'-deoxy-3'-[(O-methyl-L-tyrosyl)amino]adenosine, GLYCEROL, Peptidyl-tRNA hydrolase | | 著者 | Singh, A, Sinha, M, Bhushan, A, Kaur, P, Sharma, S, Singh, T.P. | | 登録日 | 2014-05-08 | | 公開日 | 2014-05-28 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.77 Å) | | 主引用文献 | Structural and binding studies of peptidyl-tRNA hydrolase from Pseudomonas aeruginosa provide a platform for the structure-based inhibitor design against peptidyl-tRNA hydrolase
Biochem.J., 463, 2014
|
|
4QD3
 
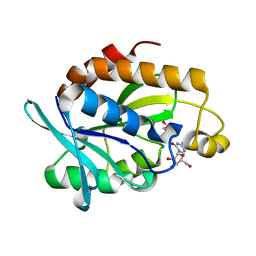 | | Crystal structure of Peptidyl-tRNA hydrolase from Pseudomonas aeruginosa with 5-azacytidine at 1.89 Angstrom resolution | | 分子名称: | 4-amino-1-(beta-D-ribofuranosyl)-1,3,5-triazin-2(1H)-one, GLYCEROL, Peptidyl-tRNA hydrolase | | 著者 | Singh, A, Gautam, L, Sinha, M, Bhushan, A, Kaur, P, Sharma, S, Singh, T.P. | | 登録日 | 2014-05-13 | | 公開日 | 2014-06-25 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.89 Å) | | 主引用文献 | Structural and binding studies of peptidyl-tRNA hydrolase from Pseudomonas aeruginosa provide a platform for the structure-based inhibitor design against peptidyl-tRNA hydrolase
Biochem.J., 463, 2014
|
|
