2EJT
 
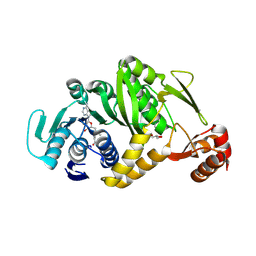 | | Complex structure of Trm1 from Pyrococcus horikoshii with S-adenosyl-L-Methionine | | Descriptor: | GLYCEROL, N(2),N(2)-dimethylguanosine tRNA methyltransferase, S-ADENOSYLMETHIONINE | | Authors: | Ihsanawati, Shirouzu, M, Bessho, Y, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-20 | | Release date: | 2007-09-25 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of tRNA N(2),N(2)-Guanosine Dimethyltransferase Trm1 from Pyrococcus horikoshii
J.Mol.Biol., 383, 2008
|
|
2E7J
 | |
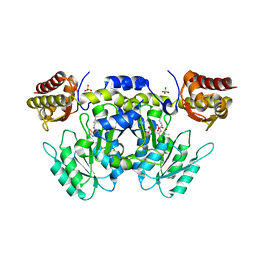
2E87
 
 | | Crystal structure of hypothetical GTP-binding protein PH1320 from Pyrococcus horikoshii OT3, in complex with GDP | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Hypothetical protein PH1320, SULFATE ION | | Authors: | Kukimoto-Niino, M, Bessho, Y, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-01-19 | | Release date: | 2007-07-24 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of hypothetical GTP-binding protein PH1320 from Pyrococcus horikoshii OT3, in complex with GDP
To be Published
|
|
2EB2
 
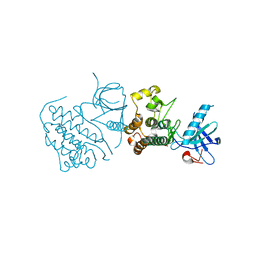 | | Crystal structure of mutated EGFR kinase domain (G719S) | | Descriptor: | Epidermal growth factor receptor | | Authors: | Yoshikawa, S, Kukimoto-Niino, M, Chen, L, Liu, Z.J, Wang, B.C, Shirouzu, M, Senba, K, Yamamoto, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-02-06 | | Release date: | 2008-02-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis for the altered drug sensitivities of non-small cell lung cancer-associated mutants of human epidermal growth factor receptor
Oncogene, 2012
|
|
2EBG
 
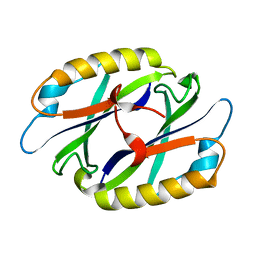 | |
2EJ7
 
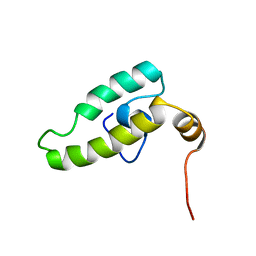 | | Solution structure of the DnaJ domain of the human protein HCG3, a hypothetical protein tmp_locus_21 | | Descriptor: | HCG3 gene | | Authors: | Ohinishi, S, Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-15 | | Release date: | 2007-09-18 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the DnaJ domain of the human protein HCG3, a hypothetical protein tmp_locus_21
To be Published
|
|
2DZK
 
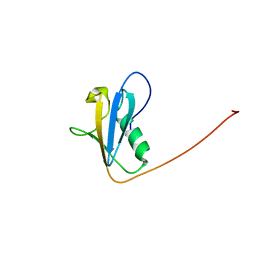 | | Structure of the UBX domain in Mouse UBX Domain-Containing Protein 2 | | Descriptor: | UBX domain-containing protein 2 | | Authors: | Zhao, C, Yoneyama, M, Koshiba, S, Watanabe, S, Harada, T, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-09-29 | | Release date: | 2007-03-29 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure of the UBX domain in Mouse UBX Domain-Containing Protein 2
To be Published
|
|
2EGI
 
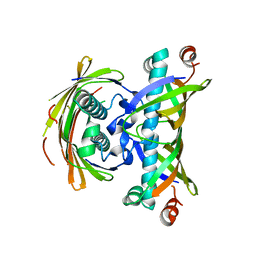 | |
2EJ8
 
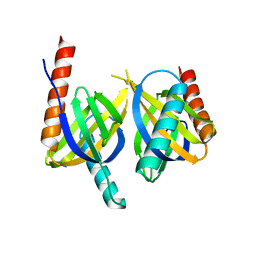 | | Crystal structure of APPL1 PTB domain at 1.8A | | Descriptor: | DCC-interacting protein 13 alpha | | Authors: | Tanabe, H, Hosaka, T, Murayama, K, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-15 | | Release date: | 2008-03-18 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal structure of APPL1 PTB domain at 1.8A
To be Published
|
|
2EJF
 
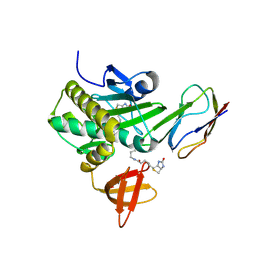 | | Crystal Structure Of The Biotin Protein Ligase (Mutations R48A and K111A) and Biotin Carboxyl Carrier Protein Complex From Pyrococcus Horikoshii OT3 | | Descriptor: | 149aa long hypothetical methylmalonyl-CoA decarboxylase gamma chain, 235aa long hypothetical biotin--[acetyl-CoA-carboxylase] ligase, ADENOSINE, ... | | Authors: | Bagautdinov, B, Matsuura, Y, Bagautdinova, S, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-16 | | Release date: | 2008-03-18 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Protein biotinylation visualized by a complex structure of biotin protein ligase with a substrate
J.Biol.Chem., 283, 2008
|
|
2EGE
 | |
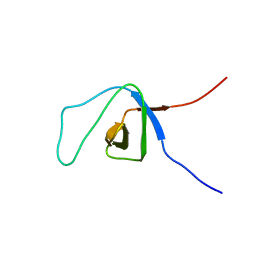
2EKB
 
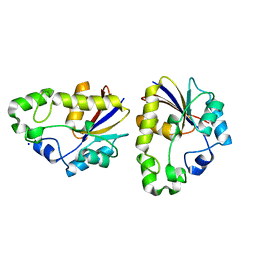 | | Structural study of Project ID TTHB049 from Thermus thermophilus HB8 (L19M) | | Descriptor: | Alpha-ribazole-5'-phosphate phosphatase, SODIUM ION | | Authors: | Asada, Y, Taketa, M, Tanaka, Y, Matsuura, Y, Kunishima, N, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-22 | | Release date: | 2007-09-25 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural study of Project ID TTHB049 from Thermus thermophilus HB8 (L19M)
To be Published
|
|
2EKP
 
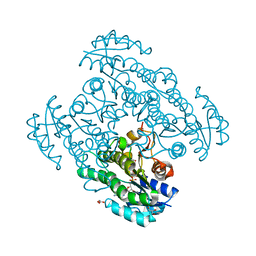 | |
2E9H
 
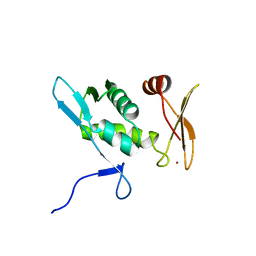 | | Solution structure of the eIF-5_eIF-2B domain from human Eukaryotic translation initiation factor 5 | | Descriptor: | Eukaryotic translation initiation factor 5, ZINC ION | | Authors: | Tomizawa, T, Tochio, N, Koshiba, S, Watanabe, S, Harada, T, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-01-25 | | Release date: | 2008-01-15 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the eIF-5_eIF-2B domain from human Eukaryotic translation initiation factor 5
To be Published
|
|
2EA9
 
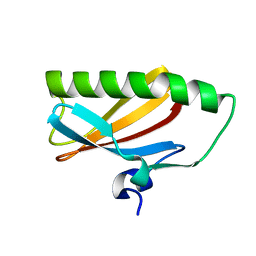 | |
2EB3
 
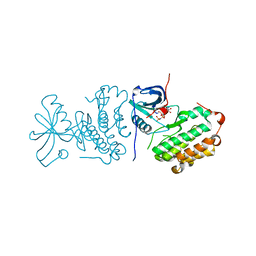 | | Crystal structure of mutated EGFR kinase domain (L858R) in complex with AMPPNP | | Descriptor: | Epidermal growth factor receptor, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Yoshikawa, S, Kukimoto-Niino, M, Shirouzu, M, Senba, K, Yamamoto, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-02-06 | | Release date: | 2008-02-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Structural basis for the altered drug sensitivities of non-small cell lung cancer-associated mutants of human epidermal growth factor receptor
Oncogene, 2012
|
|
2EHP
 
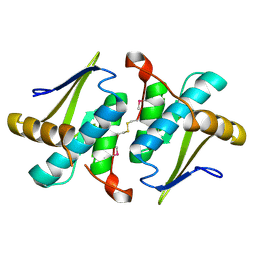 | | Crystal Structure of a Putative protein (AQ1627) from Aquifex aeolicus | | Descriptor: | aq_1627 protein | | Authors: | Kumarevel, T.S, Karthe, P, Nakano, N, Kuramitsu, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-07 | | Release date: | 2007-09-11 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structure of a Putative protein (AQ1627) from Aquifex aeolicus
To be Published
|
|
2EHW
 
 | |
2EI5
 
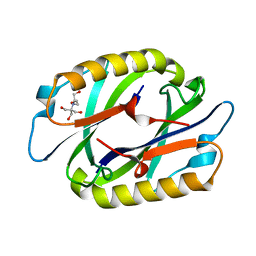 | | Crystal Structure of Hypothetical protein(TTHA0061) from Thermus thermophilus | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Hypothetical protein TTHA0061 | | Authors: | Niwa, H, Kumarevel, T.S, Kuramitsu, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-12 | | Release date: | 2007-09-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Crystal Structure of Hypothetical protein(TTHA0061) from Thermus thermophilus
To be Published
|
|
2EGJ
 
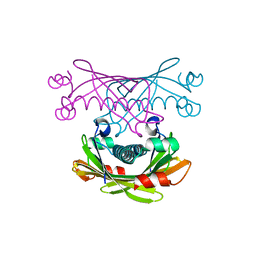 | |
2DX6
 
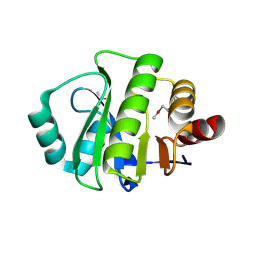 | | Crystal structure of conserved hypothetical protein, TTHA0132 from Thermus thermophilus HB8 | | Descriptor: | ACETATE ION, Hypothetical protein TTHA0132, ISOPROPYL ALCOHOL | | Authors: | Kishishita, S, Murayama, K, Terada, T, Shirouzu, M, Kuramitsu, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-08-24 | | Release date: | 2007-02-24 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Crystal structure of conserved hypothetical protein, TTHA0132 from Thermus thermophilus HB8
To be Published
|
|
2E29
 
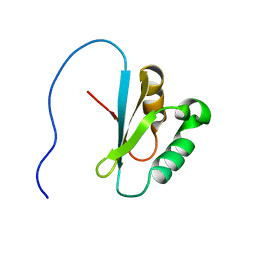 | | Solution structure of the GUCT domain from human ATP-dependent RNA helicase DDX50, DEAD box protein 50 | | Descriptor: | ATP-dependent RNA helicase DDX50 | | Authors: | Ohnishi, S, Paakkonen, K, Guntert, P, Sato, M, Koshiba, S, Harada, T, Watanabe, S, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-11-10 | | Release date: | 2007-05-15 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the GUCT domain from human RNA helicase II/Gubeta reveals the RRM fold, but implausible RNA interactions
Proteins, 74, 2008
|
|
2EJU
 
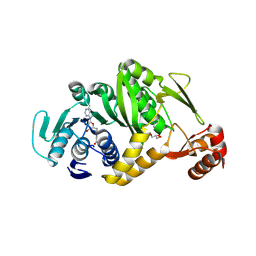 | | Complex structure of Trm1 from Pyrococcus horikoshii with S-adenosyl-L-Homocystein | | Descriptor: | GLYCEROL, N(2),N(2)-dimethylguanosine tRNA methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Ihsanawati, Shirouzu, M, Bessho, Y, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-20 | | Release date: | 2007-09-25 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of tRNA N(2),N(2)-Guanosine Dimethyltransferase Trm1 from Pyrococcus horikoshii
J.Mol.Biol., 383, 2008
|
|
2EP5
 
 | |
2EQW
 
 | | Solution structure of the 6th C2H2 type zinc finger domain of Zinc finger protein 484 | | Descriptor: | ZINC ION, Zinc finger protein 484 | | Authors: | Imai, M, Suzuki, S, Muto, Y, Inoue, M, Kigawa, T, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-30 | | Release date: | 2007-10-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the 6th C2H2 type zinc finger domain of Zinc finger protein 484
To be Published
|
|
