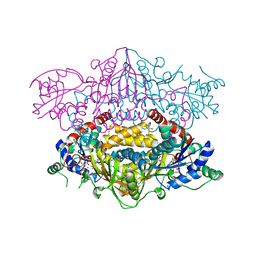
4XFJ
 
 | |
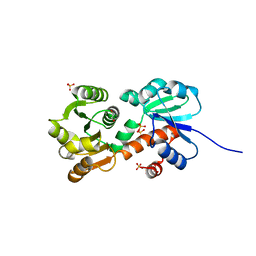
4XIJ
 
 | |
3IKF
 
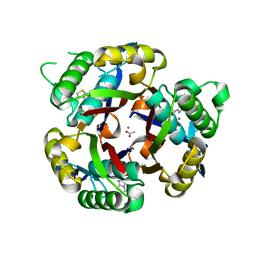 | | Crystal structure of 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase from Burkholderia pseudomallei with FOL fragment 717, imidazo[2,,1-b][1,3]thiazol-6-ylmethanol | | Descriptor: | 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase, ACETATE ION, CHLORIDE ION, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-08-05 | | Release date: | 2009-08-18 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Leveraging structure determination with fragment screening for infectious disease drug targets: MECP synthase from Burkholderia pseudomallei.
J Struct Funct Genomics, 12, 2011
|
|
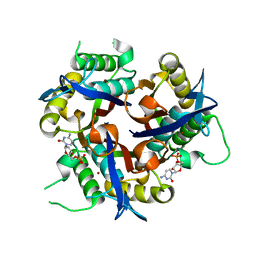
3F0G
 
 | |
3JVH
 
 | |
4E51
 
 | |
7T1W
 
 | |
7T1X
 
 | |
7SSQ
 
 | |
7SSR
 
 | | Crystal Structure of Ebola zaire Envelope glycoprotein GP in complex with compound ARN0075093 | | Descriptor: | (1R,2s,3S,5s,7s)-N-[(1r,4r)-4-(aminomethyl)cyclohexyl]-5-phenyladamantane-2-carboxamide, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2021-11-11 | | Release date: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of Ebola zaire Envelope glycoprotein GP in complex with compound ARN0075093
to be published
|
|
7TI7
 
 | |
7U5Y
 
 | |
7U5F
 
 | |
7U35
 
 | |
7U4H
 
 | |
7U56
 
 | |
7U5Q
 
 | |
7ULH
 
 | |
7ULZ
 
 | |
7UG3
 
 | |
7V0H
 | |
5SCQ
 
 | |
5SCW
 
 | |
5SD2
 
 | |
5SCU
 
 | |
