9NSC
 
 | | Bacterial Pictet-Spenglerase KslB in complex with L-Trp | | Descriptor: | Pictet-Spenglerase, TRYPTOPHAN | | Authors: | Kim, K, Kim, W. | | Deposit date: | 2025-03-16 | | Release date: | 2025-04-23 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural and mechanistic insights into KslB, a bacterial Pictet-Spenglerase in kitasetaline biosynthesis.
Rsc Chem Biol, 6, 2025
|
|
9NST
 
 | | Bacterial Pictet-Spenglerase KslB in complex with product of L-Trp and a-ketoglutaric acid | | Descriptor: | (1~{S},3~{S})-1-(3-hydroxy-3-oxopropyl)-2,3,4,9-tetrahydropyrido[3,4-b]indole-1,3-dicarboxylic acid, Pictet-Spenglerase | | Authors: | Kim, K, Kim, W. | | Deposit date: | 2025-03-17 | | Release date: | 2025-04-23 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structural and mechanistic insights into KslB, a bacterial Pictet-Spenglerase in kitasetaline biosynthesis.
Rsc Chem Biol, 6, 2025
|
|
9NSS
 
 | |
9NS6
 
 | | KslB apoenzyme | | Descriptor: | Pictet-Spenglerase | | Authors: | Kim, K, Kim, W. | | Deposit date: | 2025-03-16 | | Release date: | 2025-04-23 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural and mechanistic insights into KslB, a bacterial Pictet-Spenglerase in kitasetaline biosynthesis.
Rsc Chem Biol, 6, 2025
|
|
9NSU
 
 | | Bacterial Pictet-Spenglerase KslB in complex with product of L-Trp and succinic semialdehyde | | Descriptor: | (1S,3S)-1-(2-carboxyethyl)-2,3,4,9-tetrahydro-1H-pyrido[3,4-b]indole-3-carboxylic acid, Pictet-Spenglerase | | Authors: | Kim, K, Kim, W. | | Deposit date: | 2025-03-17 | | Release date: | 2025-04-23 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Structural and mechanistic insights into KslB, a bacterial Pictet-Spenglerase in kitasetaline biosynthesis.
Rsc Chem Biol, 6, 2025
|
|
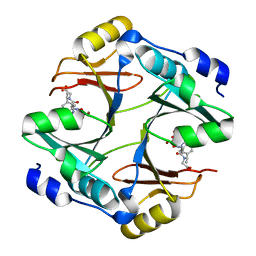
4OPN
 
 | |
6LAT
 
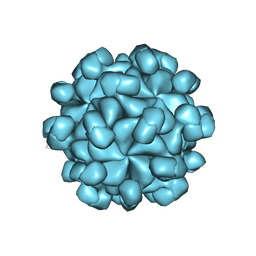 | | The cryo-EM structure of HEV VLP | | Descriptor: | Protein ORF2 | | Authors: | Zheng, Q, He, M, Li, S. | | Deposit date: | 2019-11-13 | | Release date: | 2019-12-04 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Viral neutralization by antibody-imposed physical disruption.
Proc.Natl.Acad.Sci.USA, 2019
|
|
6PKA
 | |
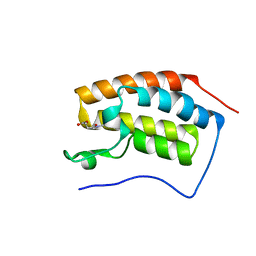
6PRT
 
 | |
6PS9
 
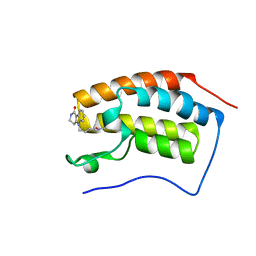 | | Crystal structure of BRD4 bromodomain 1 with N-methylpyrrolidin-2-one (NMP) derivative 17 (5-{2-[(3R)-1-methyl-5-oxopyrrolidin-3-yl]ethyl}-2,3,4,5-tetrahydro-1H-pyrido[4,3-b]indol-1-one) | | Descriptor: | 5-{2-[(3R)-1-methyl-5-oxopyrrolidin-3-yl]ethyl}-2,3,4,5-tetrahydro-1H-pyrido[4,3-b]indol-1-one, Bromodomain-containing protein 4 | | Authors: | Ilyichova, O.V, Scanlon, M.J. | | Deposit date: | 2019-07-12 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.21 Å) | | Cite: | Synthesis and elaboration of N-methylpyrrolidone as an acetamide fragment substitute in bromodomain inhibition.
Bioorg.Med.Chem., 27, 2019
|
|
6PSB
 
 | | Crystal structure of BRD4 bromodomain 1 with N-methylpyrrolidin-2-one (NMP) derivative 18 (5-{[(3R)-1-methyl-5-oxopyrrolidin-3-yl]methyl}-2,3,4,5-tetrahydro-1H-pyrido[4,3-b]indol-1-one) | | Descriptor: | 5-{[(3R)-1-methyl-5-oxopyrrolidin-3-yl]methyl}-2,3,4,5-tetrahydro-1H-pyrido[4,3-b]indol-1-one, Bromodomain-containing protein 4 | | Authors: | Ilyichova, O.V, Scanlon, M.J. | | Deposit date: | 2019-07-12 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Synthesis and elaboration of N-methylpyrrolidone as an acetamide fragment substitute in bromodomain inhibition.
Bioorg.Med.Chem., 27, 2019
|
|
6PMD
 | |
7SCP
 
 | | The crystal structure of ScoE in complex with intermediate | | Descriptor: | (3R)-3-(oxaloamino)butanoic acid, 1,2-ETHANEDIOL, FE (II) ION, ... | | Authors: | Cha, L, Chen, J, Zhou, J, Chang, W. | | Deposit date: | 2021-09-28 | | Release date: | 2022-03-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Deciphering the Reaction Pathway of Mononuclear Iron Enzyme-Catalyzed N-C Triple Bond Formation in Isocyanide Lipopeptide and Polyketide Biosynthesis
Acs Catalysis, 12, 2022
|
|
3LKJ
 
 | | Small Molecule Inhibition of the TNF Family Cyokine CD40 Ligand Through a Subunit Fracture Mechanism | | Descriptor: | (2R)-{[(2'-[(biphenyl-3-ylmethyl)carbamoyl]-6'-{[(2R)-2-(pyrrolidin-1-ylmethyl)pyrrolidin-1-yl]carbonyl}-6-{[(2R)-2-(1H-pyrrol-1-ylmethyl)pyrrolidin-1-yl]carbonyl}-4,4'-bipyridin-2-yl)carbonyl]amino}(cyclohexyl)ethanoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, CD40 ligand | | Authors: | Silvian, L.F, Whitty, A. | | Deposit date: | 2010-01-27 | | Release date: | 2011-02-02 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Small Molecule Inhibition of the TNF Family Cytokine CD40 Ligand through a Subunit Fracture Mechanism.
Acs Chem.Biol., 6, 2011
|
|
3MQ0
 
 | | Crystal Structure of Agobacterium tumefaciens repressor BlcR | | Descriptor: | Transcriptional repressor of the blcABC operon | | Authors: | Chen, L. | | Deposit date: | 2010-04-27 | | Release date: | 2011-03-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.793 Å) | | Cite: | The Agrobacterium tumefaciens Transcription Factor BlcR Is Regulated via Oligomerization.
J.Biol.Chem., 286, 2011
|
|
6VUB
 
 | |
6VUF
 
 | |
6VUJ
 
 | | Crystal structure of BRD4 bromodomain 1 with N-methylpyrrolidin-2-one (NMP) derivative 15c (N,N-diethyl-3',4'-dimethoxy-6-(1-methyl-5-oxopyrrolidin-3-yl)-[1,1'-biphenyl]-3-sulfonamide) | | Descriptor: | Bromodomain-containing protein 4, N,N-diethyl-3',4'-dimethoxy-6-[(3S)-1-methyl-5-oxopyrrolidin-3-yl][1,1'-biphenyl]-3-sulfonamide, NITRATE ION | | Authors: | Ilyichova, O.V, Scanlon, M.J, Thompson, P.E. | | Deposit date: | 2020-02-15 | | Release date: | 2020-02-26 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Substituted 1-methyl-4-phenylpyrrolidin-2-ones - Fragment-based design of N-methylpyrrolidone-derived bromodomain inhibitors.
Eur.J.Med.Chem., 191, 2020
|
|
6VUC
 
 | |
1RCS
 
 | | NMR STUDY OF TRP REPRESSOR-OPERATOR DNA COMPLEX | | Descriptor: | DNA (5'-D(*CP*GP*TP*AP*CP*TP*AP*GP*TP*TP*AP*AP*CP*TP*AP*GP*TP*AP*CP*G)-3'), TRP REPRESSOR, TRYPTOPHAN | | Authors: | Zhao, D, Zheng, Z. | | Deposit date: | 1995-05-12 | | Release date: | 1996-06-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The solution structures of the trp repressor-operator DNA complex.
J.Mol.Biol., 238, 1994
|
|
6CLV
 
 | | Staphylococcus aureus Dihydropteroate Synthase (saDHPS) F17L E208K double mutant structure | | Descriptor: | 4-{[(2-amino-4-oxo-3,4-dihydropteridin-6-yl)methyl]amino}-N-(3,4-dimethyl-1,2-oxazol-5-yl)benzene-1-sulfonamide, Dihydropteroate synthase | | Authors: | Gajewski, S, Griffith, E.C, Wu, Y, White, S.W. | | Deposit date: | 2018-03-02 | | Release date: | 2018-08-22 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Structural and Functional Basis for Recurring Sulfa Drug Resistance Mutations inStaphylococcus aureusDihydropteroate Synthase.
Front Microbiol, 9, 2018
|
|
4K3J
 
 | | Crystal structure of Onartuzumab Fab in complex with MET and HGF-beta | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Hepatocyte growth factor, Hepatocyte growth factor beta chain, ... | | Authors: | Ma, X, Starovasnik, M.A. | | Deposit date: | 2013-04-10 | | Release date: | 2013-08-28 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Monovalent antibody design and mechanism of action of onartuzumab, a MET antagonist with anti-tumor activity as a therapeutic agent.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
6CLU
 
 | |
5VZ2
 | |
5W18
 | | Staphylococcus aureus ClpP in complex with (S)-N-((2R,6S,8aS,14aS,20S,23aS)-2,6-dimethyl-5,8,14,19,23-pentaoxooctadecahydro-1H,5H,14H,19H-pyrido[2,1-i]dipyrrolo[2,1-c:2',1'-l][1]oxa[4,7,10,13]tetraazacyclohexadecin-20-yl)-3-phenyl-2-(3-phenylureido)propanamide | | Descriptor: | 9V7-PHE-SER-PRO-YCP-ALA-MP8, ATP-dependent Clp protease proteolytic subunit | | Authors: | Lee, R.E, Griffith, E.C. | | Deposit date: | 2017-06-02 | | Release date: | 2017-08-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Ureadepsipeptides as ClpP Activators.
Acs Infect Dis., 2019
|
|
