7LHN
 | |
7LHM
 | |
7LHO
 | |
7MS0
 
 | | Crystal structure of native Cg10062 | | Descriptor: | 4-oxalocrotonate tautomerase | | Authors: | Nayebi, G.H, Geiger, J.H, Draths, K. | | Deposit date: | 2021-05-10 | | Release date: | 2022-02-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | Cg10062 Catalysis Forges a Link between Acetylenecarboxylic Acid and Bacterial Metabolism.
Biochemistry, 60, 2021
|
|
7MS3
 
 | |
7MS8
 
 | |
7MS1
 
 | |
4EFG
 | |
4EXZ
 | |
4EDE
 | |
4EEJ
 | |
1P1H
 
 | | Crystal structure of the 1L-myo-inositol/NAD+ complex | | Descriptor: | Inositol-3-phosphate synthase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Jin, X, Geiger, J.H. | | Deposit date: | 2003-04-12 | | Release date: | 2003-07-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structures of NAD(+)- and NADH-bound 1-l-myo-inositol 1-phosphate synthase.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1P1K
 
 | |
1P1J
 
 | | Crystal structure of the 1L-myo-inositol 1-phosphate synthase complexed with NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, GLYCEROL, Inositol-3-phosphate synthase, ... | | Authors: | Jin, X, Geiger, J.H. | | Deposit date: | 2003-04-12 | | Release date: | 2003-07-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of NAD(+)- and NADH-bound 1-l-myo-inositol 1-phosphate synthase.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
1P1I
 
 | |
4ZH6
 
 | |
4ZH9
 
 | |
4ZR2
 
 | |
1P1F
 
 | |
3F8A
 
 | | Crystal Structure of the R132K:R111L:L121E:R59W Mutant of Cellular Retinoic Acid-Binding Protein Type II Complexed with C15-aldehyde (a retinal analog) at 1.95 Angstrom resolution. | | Descriptor: | 1,3,3-trimethyl-2-[(1E,3E)-3-methylpenta-1,3-dien-1-yl]cyclohexene, 2-[3-(2-HYDROXY-1,1-DIHYDROXYMETHYL-ETHYLAMINO)-PROPYLAMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Cellular retinoic acid-binding protein 2 | | Authors: | Jia, X, Geiger, J.H. | | Deposit date: | 2008-11-12 | | Release date: | 2009-11-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Probing Wavelength Regulation with an Engineered Rhodopsin Mimic and a C15-Retinal Analogue
Chempluschem, 77, 2012
|
|
3FA6
 | |
3CR6
 | |
3F9D
 
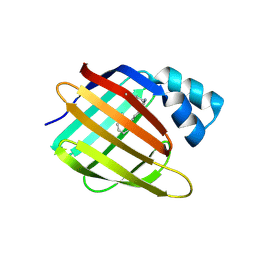 | |
1IG7
 
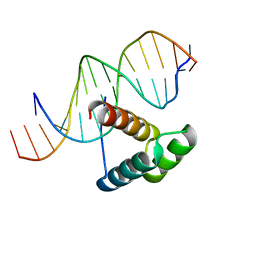 | | Msx-1 Homeodomain/DNA Complex Structure | | Descriptor: | 5'-D(*CP*AP*CP*TP*AP*AP*TP*TP*GP*AP*AP*GP*G)-3', 5'-D(P*TP*CP*CP*TP*TP*CP*AP*AP*TP*TP*AP*GP*TP*GP*AP*C)-3', Homeotic protein Msx-1 | | Authors: | Hovde, S, Abate-Shen, C, Geiger, J.H. | | Deposit date: | 2001-04-17 | | Release date: | 2001-04-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the Msx-1 homeodomain/DNA complex
Biochemistry, 40, 2001
|
|
1KI0
 
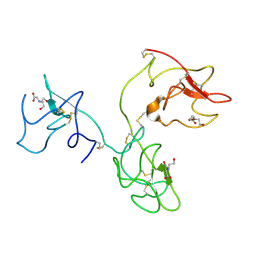 | | The X-ray Structure of Human Angiostatin | | Descriptor: | ANGIOSTATIN, BICINE | | Authors: | Abad, M.C, Arni, R.K, Grella, D.K, Castellino, F.J, Tulinsky, A, Geiger, J.H. | | Deposit date: | 2001-12-02 | | Release date: | 2002-05-29 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The X-ray crystallographic structure of the angiogenesis inhibitor angiostatin.
J.Mol.Biol., 318, 2002
|
|
