5WZ4
 
 | |
6F2P
 
 | | Structure of Paenibacillus xanthan lyase to 2.6 A resolution | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ... | | Authors: | Lo Leggio, L, Kadziola, A. | | Deposit date: | 2017-11-27 | | Release date: | 2018-12-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure and Dynamics of a Promiscuous Xanthan Lyase from Paenibacillus nanensis and the Design of Variants with Increased Stability and Activity.
Cell Chem Biol, 26, 2019
|
|
7YRT
 | |
8H5V
 
 | | Crystal structure of the FleQ domain of Vibrio cholerae FlrA | | Descriptor: | 1,2-ETHANEDIOL, SULFATE ION, flagellar regulatory protein A | | Authors: | Dasgupta, J, Chakraborty, S. | | Deposit date: | 2022-10-14 | | Release date: | 2022-11-02 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The N-terminal FleQ domain of the Vibrio cholerae flagellar master regulator FlrA plays pivotal structural roles in stabilizing its active state.
Febs Lett., 597, 2023
|
|
8SKQ
 
 | |
6A7V
 
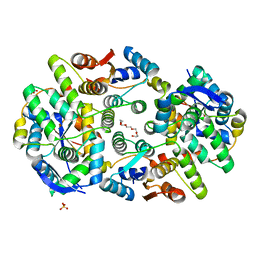 | | Crystal structure of Mycobacterium tuberculosis VapBC11 toxin-antitoxin complex | | Descriptor: | Antitoxin VapB11, PENTAETHYLENE GLYCOL, Ribonuclease VapC11, ... | | Authors: | Deep, A, Thakur, K.G. | | Deposit date: | 2018-07-04 | | Release date: | 2018-10-10 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Structural, functional and biological insights into the role of Mycobacterium tuberculosis VapBC11 toxin-antitoxin system: targeting a tRNase to tackle mycobacterial adaptation.
Nucleic Acids Res., 46, 2018
|
|
5YWR
 
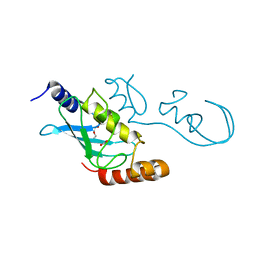 | | Crystal Structure of RING E3 ligase ZNRF1 in complex with Ube2N (Ubc13) | | Descriptor: | E3 ubiquitin-protein ligase ZNRF1, FORMIC ACID, TRIETHYLENE GLYCOL, ... | | Authors: | Behera, A.P, Naskar, P, Datta, A.B. | | Deposit date: | 2017-11-30 | | Release date: | 2018-06-06 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Structural insights into the nanomolar affinity of RING E3 ligase ZNRF1 for Ube2N and its functional implications.
Biochem. J., 475, 2018
|
|
5BKI
 
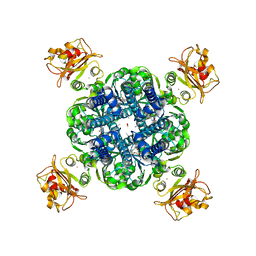 | | Blocker-free closed MthK channel in nanodisc | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, CADMIUM ION, Calcium-gated potassium channel MthK, ... | | Authors: | Fan, C, Nimigean, C.M. | | Deposit date: | 2021-03-19 | | Release date: | 2022-09-21 | | Last modified: | 2023-10-04 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Calcium-gated potassium channel blockade via membrane-facing fenestrations.
Nat.Chem.Biol., 2023
|
|
5BKK
 
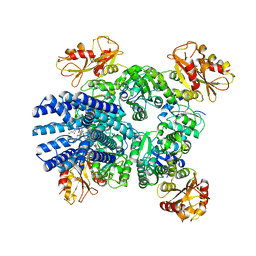 | | bbTBA-bound closed MthK channel in nanodisc | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, (~{R})-phenyl-[4-[(tributyl-$l^{4}-azanyl)methyl]phenyl]methanol, Calcium-gated potassium channel MthK, ... | | Authors: | Fan, C, Nimigean, C.M. | | Deposit date: | 2021-03-19 | | Release date: | 2022-09-21 | | Last modified: | 2023-10-04 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Calcium-gated potassium channel blockade via membrane-facing fenestrations.
Nat.Chem.Biol., 2023
|
|
5BKJ
 
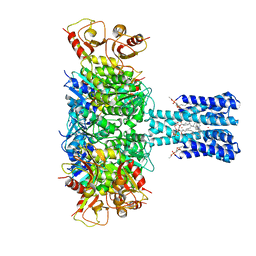 | | TPeA-bound closed MthK channel in nanodisc | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, 1-(tripentyl-$l^{4}-azanyl)pentane, Calcium-gated potassium channel MthK, ... | | Authors: | Fan, C, Nimigean, C.M. | | Deposit date: | 2021-03-19 | | Release date: | 2022-09-21 | | Last modified: | 2023-10-04 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Calcium-gated potassium channel blockade via membrane-facing fenestrations.
Nat.Chem.Biol., 2023
|
|
7LY1
 
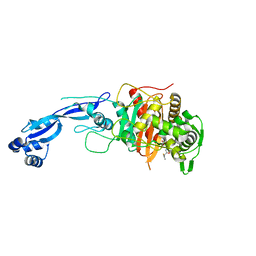 | |
2HC8
 
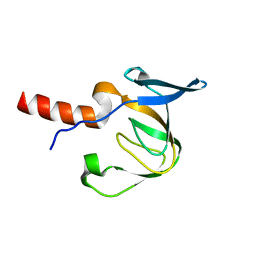 | | Structure of the A. fulgidus CopA A-domain | | Descriptor: | Cation-transporting ATPase, P-type | | Authors: | Sazinsky, M.H, Agawal, S, Arguello, J.M, Rosenzweig, A.C. | | Deposit date: | 2006-06-15 | | Release date: | 2006-09-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure of the Actuator Domain from the Archaeoglobus fulgidus Cu(+)-ATPase(,).
Biochemistry, 45, 2006
|
|
2FEA
 | |
1VRM
 
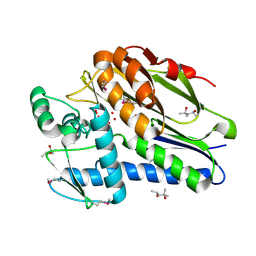 | |
1VR0
 
 | |
1VQ3
 | |
1VR8
 
 | |
1ZX8
 
 | |
2FNO
 
 | |
2GHR
 
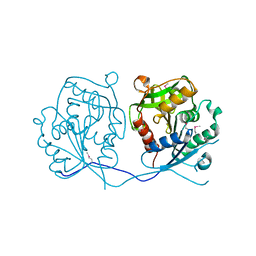 | |
3N0C
 
 | | TM0449 mutant crystal grown by hanging drop method | | Descriptor: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, FLAVIN-ADENINE DINUCLEOTIDE, Thymidylate synthase thyX | | Authors: | Mathews, I.I. | | Deposit date: | 2010-05-13 | | Release date: | 2011-05-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Diffraction study of protein crystals grown in cryoloops and micromounts.
J.Appl.Crystallogr., 43, 2010
|
|
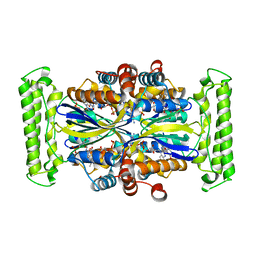
3N0B
 
 | | TM0449 mutant crystals grown in loops/micromounts | | Descriptor: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, FLAVIN-ADENINE DINUCLEOTIDE, Thymidylate synthase thyX | | Authors: | Mathews, I.I. | | Deposit date: | 2010-05-13 | | Release date: | 2011-05-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Diffraction study of protein crystals grown in cryoloops and micromounts.
J.Appl.Crystallogr., 43, 2010
|
|
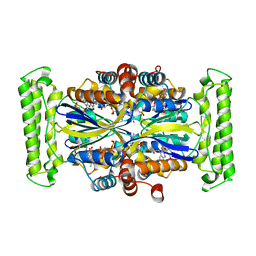
1VR3
 | |
1VQ0
 
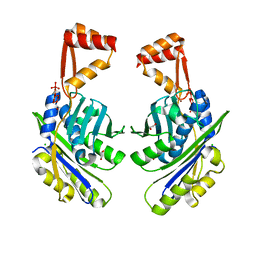 | |
1VQR
 
 | |
