8AKI
 
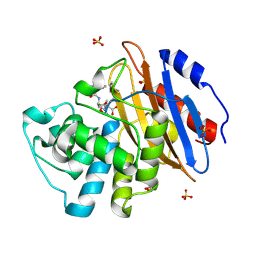 | | Acyl-enzyme complex of ampicillin bound to deacylation mutant KPC-2 (E166Q) | | Descriptor: | (2R,4S)-2-[(1R)-1-{[(2R)-2-amino-2-phenylacetyl]amino}-2-oxoethyl]-5,5-dimethyl-1,3-thiazolidine-4-carboxylic acid, Carbapenem-hydrolyzing beta-lactamase KPC, GLYCEROL, ... | | Authors: | Tooke, C.L, Hinchliffe, P, Spencer, J. | | Deposit date: | 2022-07-29 | | Release date: | 2023-03-08 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Tautomer-Specific Deacylation and Omega-Loop Flexibility Explain the Carbapenem-Hydrolyzing Broad-Spectrum Activity of the KPC-2 beta-Lactamase.
J.Am.Chem.Soc., 145, 2023
|
|
9BA2
 
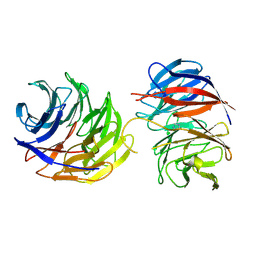 | | Crystal structure of the binary complex of DCAF1 and WDR5 | | Descriptor: | DDB1- and CUL4-associated factor 1, IMIDAZOLE, WD repeat-containing protein 5 | | Authors: | Mabanglo, M.F, Wilson, B.J, Srivastava, S, Al-awar, R, Vedadi, M. | | Deposit date: | 2024-04-03 | | Release date: | 2024-11-06 | | Last modified: | 2024-12-04 | | Method: | X-RAY DIFFRACTION (2.97 Å) | | Cite: | Crystal structures of DCAF1-PROTAC-WDR5 ternary complexes provide insight into DCAF1 substrate specificity.
Nat Commun, 15, 2024
|
|
9BD5
 
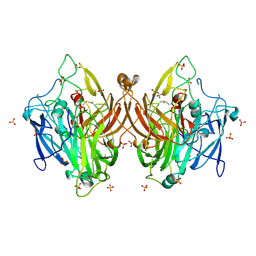 | | Laccase from Bacillus licheniformis | | Descriptor: | COPPER (II) ION, SULFATE ION, Spore coat protein A | | Authors: | Habib, M.H, Smith, T.J. | | Deposit date: | 2024-04-11 | | Release date: | 2024-11-20 | | Last modified: | 2024-12-18 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Polymerization potential of a bacterial CotA-laccase for beta-naphthol: enzyme structure and comprehensive polymer characterization.
Front Microbiol, 15, 2024
|
|
9B8J
 
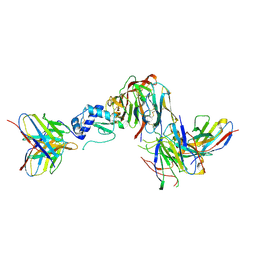 | | GP38-GnH-DS-Gc in the pre-fusion conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ADI-36125 heavy chain, ADI-36125 light chain, ... | | Authors: | McFadden, E, McLellan, J.S. | | Deposit date: | 2024-03-30 | | Release date: | 2024-12-11 | | Last modified: | 2025-02-05 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Engineering and structures of Crimean-Congo hemorrhagic fever virus glycoprotein complexes.
Cell, 188, 2025
|
|
9C0O
 
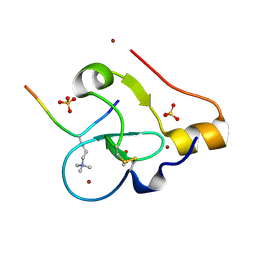 | |
9CS7
 
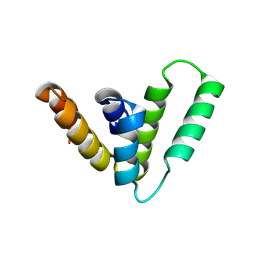 | | Structure of Azospirillum bacterial CARD crystal form 1 | | Descriptor: | Serine protease | | Authors: | Wein, T, Millman, A, Lange, K, Yirmiya, E, Hadary, R, Garb, J, Melamed, S, Steinruecke, F, HIll, A.B, Kranzusch, P.J, Sorek, R. | | Deposit date: | 2024-07-23 | | Release date: | 2025-01-29 | | Last modified: | 2025-03-26 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | CARD domains mediate anti-phage defence in bacterial gasdermin systems.
Nature, 639, 2025
|
|
9CS8
 
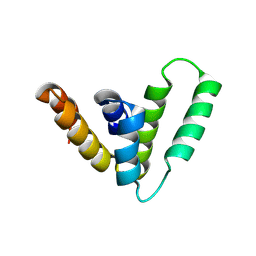 | | Structure of Azospirillum bacterial CARD crystal form 2 | | Descriptor: | Serine protease | | Authors: | Wein, T, Millman, A, Lange, K, Yirmiya, E, Hadary, R, Garb, J, Melamed, S, Steinruecke, F, Hill, A.B, Kranzusch, P.J, Sorek, R. | | Deposit date: | 2024-07-23 | | Release date: | 2025-01-29 | | Last modified: | 2025-03-26 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | CARD domains mediate anti-phage defence in bacterial gasdermin systems.
Nature, 639, 2025
|
|
9CSH
 
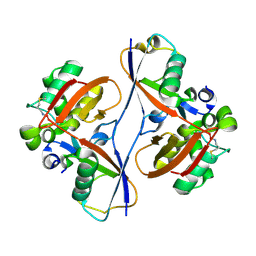 | |
9C3G
 
 | |
9CYB
 
 | | SARS-CoV-2 PLpro in complex with inhibitor WEHI-P1 | | Descriptor: | Papain-like protease, SUCCINIC ACID, [(3R)-1-cyclopentylpiperidin-3-yl](6-methoxynaphthalen-2-yl)methanone | | Authors: | Calleja, D.J, Lechtenberg, B.C, Kuchel, N.W, Devine, S.M, Bader, S.M, Doerflinger, M, Mitchell, J.P, Lessene, G, Komander, D. | | Deposit date: | 2024-08-01 | | Release date: | 2025-04-09 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | A novel PLpro inhibitor improves outcomes in a pre-clinical model of long COVID.
Nat Commun, 16, 2025
|
|
9CYD
 
 | | SARS-CoV-2 PLpro in complex with inhibitor WEHI-P4 | | Descriptor: | (1S,4s)-4-{(3R)-3-[(E)-(methoxyimino)(6-methoxynaphthalen-2-yl)methyl]piperidin-1-yl}cyclohexan-1-ol, CHLORIDE ION, Papain-like protease, ... | | Authors: | Calleja, D.J, Lechtenberg, B.C, Kuchel, N.W, Devine, S.M, Bader, S.M, Doerflinger, M, Mitchell, J.P, Lessene, G, Komander, D. | | Deposit date: | 2024-08-02 | | Release date: | 2025-04-09 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A novel PLpro inhibitor improves outcomes in a pre-clinical model of long COVID.
Nat Commun, 16, 2025
|
|
9CYK
 
 | | SARS-CoV-2 PLpro in complex with inhibitor WEHI-P24 | | Descriptor: | ACETIC ACID, GLYCEROL, Papain-like protease, ... | | Authors: | Calleja, D.J, Lechtenberg, B.C, Kuchel, N.W, Devine, S.M, Bader, S.M, Doerflinger, M, Mitchell, J.P, Lessene, G, Komander, D. | | Deposit date: | 2024-08-02 | | Release date: | 2025-04-09 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | A novel PLpro inhibitor improves outcomes in a pre-clinical model of long COVID.
Nat Commun, 16, 2025
|
|
9CYC
 
 | | SARS-CoV-2 PLpro in complex with inhibitor WEHI-P2 | | Descriptor: | (E)-1-[(3R)-1-cyclopentylpiperidin-3-yl]-N-methoxy-1-(6-methoxynaphthalen-2-yl)methanimine, ACETIC ACID, GLYCEROL, ... | | Authors: | Calleja, D.J, Lechtenberg, B.C, Kuchel, N.W, Devine, S.M, Bader, S.M, Doerflinger, M, Mitchel, J.P, Lessene, G, Komander, D. | | Deposit date: | 2024-08-01 | | Release date: | 2025-04-09 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | A novel PLpro inhibitor improves outcomes in a pre-clinical model of long COVID.
Nat Commun, 16, 2025
|
|
9C97
 
 | |
9CMR
 
 | | Saccharomyces cerevisiae Tom1 HECT domain | | Descriptor: | E3 ubiquitin-protein ligase TOM1, SULFATE ION | | Authors: | Schubert, H.L, Hill, C.P. | | Deposit date: | 2024-07-15 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Tom1p ubiquitin ligase structure, interaction with Spt6p, and function in maintaining normal transcript levels and the stability of chromatin in promoters.
To Be Published
|
|
9E3B
 
 | | Cryo-EM structure of PRMT5/WDR77 in complex with 6S complex | | Descriptor: | Methylosome protein WDR77, Methylosome subunit pICln, Protein arginine N-methyltransferase 5, ... | | Authors: | Jin, C.Y, Hunkeler, M, Fischer, E.S. | | Deposit date: | 2024-10-23 | | Release date: | 2025-02-19 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Substrate adaptors are flexible tethering modules that enhance substrate methylation by the arginine methyltransferase PRMT5.
J.Biol.Chem., 301, 2025
|
|
9DGZ
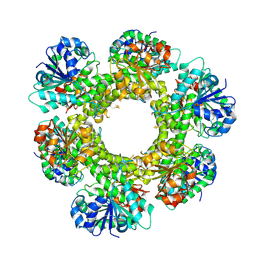 
 | |
9DH0
 
 | | The Cryo-EM structure of recombinantly expressed hUGDH in complex with UDP-4-keto-xylose | | Descriptor: | (2R,3R,4R)-3,4-dihydroxy-5-oxooxan-2-yl [(2R,3S,4R,5R)-5-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)-3,4-dihydroxyoxolan-2-yl]methyl dihydrogen diphosphate (non-preferred name), UDP-glucose 6-dehydrogenase | | Authors: | Kadirvelraj, R, Walsh Jr, R.M, Wood, Z.W. | | Deposit date: | 2024-09-03 | | Release date: | 2025-03-05 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (2.38 Å) | | Cite: | Cryo-EM Structure of Recombinantly Expressed hUGDH Unveils a Hidden, Alternative Allosteric Inhibitor.
Biochemistry, 64, 2025
|
|
9EGW
 
 | |
9EGV
 
 | | HOIL-1 RING2 domain bound to ubiquitin | | Descriptor: | CHLORIDE ION, Polyubiquitin-C, RanBP-type and C3HC4-type zinc finger-containing protein 1, ... | | Authors: | Wang, X.S, Lechtenberg, B.C. | | Deposit date: | 2024-11-21 | | Release date: | 2024-12-11 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The RBR E3 ubiquitin ligase HOIL-1 can ubiquitinate diverse non-protein substrates in vitro.
Life Sci Alliance, 8, 2025
|
|
9EC2
 
 | | Crystal structure of SAMHD1 dimer bound to an inhibitor obtained from high-throughput chemical tethering to the guanine antiviral acyclovir | | Descriptor: | Deoxynucleoside triphosphate triphosphohydrolase SAMHD1, FE (III) ION, N-[5-({2-[(2-amino-6-oxo-1,6-dihydro-9H-purin-9-yl)methoxy]ethyl}amino)-5-oxopentyl]-4,7-dibromo-3-hydroxynaphthalene-2-carboxamide | | Authors: | Egleston, M, Dong, L, Howlader, A.H, Bhat, S, Orris, B, Lopez-Rovira, L.M, Bianchet, M.A, Greenberg, M.M, Stivers, J.T. | | Deposit date: | 2024-11-13 | | Release date: | 2025-03-12 | | Last modified: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Inhibitors of SAMHD1 Obtained from Chemical Tethering to the Guanine Antiviral Acyclovir.
Biochemistry, 64, 2025
|
|
9DYB
 
 | |
9DYA
 
 | | CHIP-TPR in complex with the Hsp70 tail | | Descriptor: | E3 ubiquitin-protein ligase CHIP, Heat shock 70 kDa protein 1A, SULFATE ION | | Authors: | Zhang, H, Nix, J.C, Page, R.C. | | Deposit date: | 2024-10-13 | | Release date: | 2025-03-26 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Phosphorylation-State Modulated Binding of HSP70: Structural Insights and Compensatory Protein Engineering.
Biorxiv, 2025
|
|
9DJT
 
 | | Ternary complex structure of Cereblon-DDB1 bound to WIZ(ZF7) and the molecular glue WIZ-5 | | Descriptor: | (3S)-3-(5-{[(4R)-6-ethyl-6-azaspiro[2.5]octan-4-yl]oxy}-1-oxo-1,3-dihydro-2H-isoindol-2-yl)piperidine-2,6-dione, DNA damage-binding protein 1, Protein Wiz, ... | | Authors: | Partridge, J.R, Ma, X, Ornelas, E. | | Deposit date: | 2024-09-06 | | Release date: | 2024-11-27 | | Last modified: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Discovery and Optimization of First-in-Class Molecular Glue Degraders of the WIZ Transcription Factor for Fetal Hemoglobin Induction to Treat Sickle Cell Disease.
J.Med.Chem., 67, 2024
|
|
9E6S
 
 | |
