2BV3
 
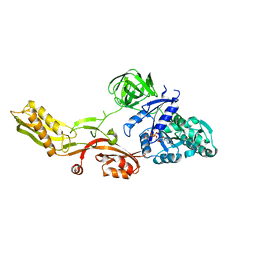 | | Crystal structure of a mutant elongation factor G trapped with a GTP analogue | | Descriptor: | ELONGATION FACTOR G, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER | | Authors: | Hansson, S, Singh, R, Gudkov, A.T, Liljas, A, Logan, D.T. | | Deposit date: | 2005-06-22 | | Release date: | 2005-08-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of a Mutant Elongation Factor G Trapped with a GTP Analogue.
FEBS Lett., 579, 2005
|
|
8P27
 
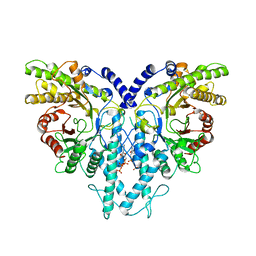 | | Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, dATP-bound state | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, Anaerobic ribonucleoside-triphosphate reductase, MAGNESIUM ION | | Authors: | Banerjee, I, Bimai, O, Sjoberg, B.M, Logan, D.T. | | Deposit date: | 2023-05-15 | | Release date: | 2023-08-30 | | Last modified: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (2.73 Å) | | Cite: | Nucleotide binding to the ATP-cone in anaerobic ribonucleotide reductases allosterically regulates activity by modulating substrate binding.
Elife, 12, 2024
|
|
8P23
 
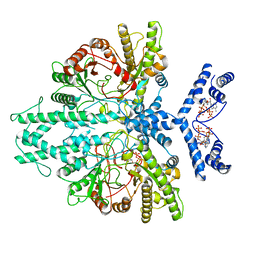 | | Cryo-EM structure of the anaerobic ribonucleotide reductase from Prevotella copri in its dimeric, ATP/CTP-bound state | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Anaerobic ribonucleoside-triphosphate reductase, CYTIDINE-5'-TRIPHOSPHATE, ... | | Authors: | Banerjee, I, Bimai, O, Sjoberg, B.M, Logan, D.T. | | Deposit date: | 2023-05-14 | | Release date: | 2023-08-30 | | Last modified: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (3.17 Å) | | Cite: | Nucleotide binding to the ATP-cone in anaerobic ribonucleotide reductases allosterically regulates activity by modulating substrate binding.
Elife, 12, 2024
|
|
4XCR
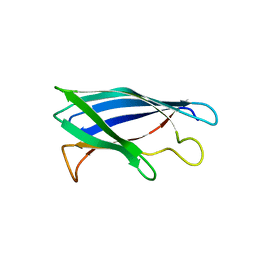 
 | | Monomeric Human Cu,Zn Superoxide dismutase, loops IV and VII deleted, apo form, mutant I35A | | Descriptor: | Superoxide dismutase [Cu-Zn] | | Authors: | Wang, H, Logan, D.T, Danielsson, J, Mu, X, Binolfi, A, Theillet, F, Bekei, B, Lang, L, Wennerstrom, H, Selenko, P, Oliveberg, M. | | Deposit date: | 2014-12-18 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.602 Å) | | Cite: | Thermodynamics of protein destabilization in live cells.
Proc. Natl. Acad. Sci. U.S.A., 112, 2015
|
|
7TPS
 
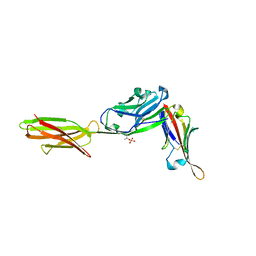 | | Crystal structure of ALPN-202 (engineered CD80 vIgD) in complex with PD-L1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Demonte, D.W, Maurer, M.F, Akutsu, M, Kimbung, Y.R, Logan, D.T, Walse, B. | | Deposit date: | 2022-01-26 | | Release date: | 2022-03-16 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | The engineered CD80 variant fusion therapeutic davoceticept combines checkpoint antagonism with conditional CD28 costimulation for anti-tumor immunity.
Nat Commun, 13, 2022
|
|
9HE8
 
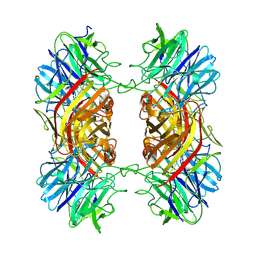 | |
9FXK
 
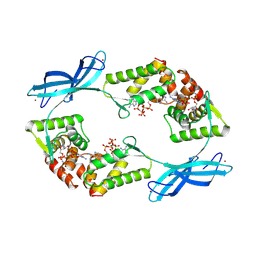 | |
9FVR
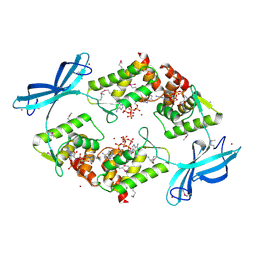 
 | | Transcription repressor NrdR from E. coli, ATP/dATP-bound state, SeMet protein | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Transcriptional repressor NrdR, ... | | Authors: | Bimai, O, Sjoberg, B.M, Rozman Grinberg, I, Logan, D.T. | | Deposit date: | 2024-06-28 | | Release date: | 2025-03-19 | | Last modified: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (3.099 Å) | | Cite: | Bacterial transcriptional repressor NrdR - a flexible multifactorial nucleotide sensor.
Febs J., 292, 2025
|
|
9FYJ
 
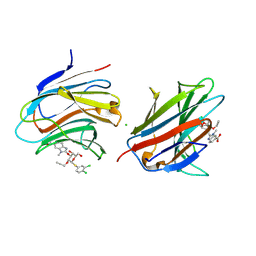 | | N-terminal domain of human galectin-8 in complex with an alpha-galactoside ligand | | Descriptor: | 2-[[(2~{R},3~{R},4~{S},5~{S},6~{R})-2-(3,4-dichlorophenyl)sulfanyl-6-(hydroxymethyl)-5-oxidanyl-3-prop-2-ynoxy-oxan-4-yl]oxymethyl]-3-methyl-benzimidazole-5-carboxylic acid, CHLORIDE ION, Isoform 1 of Galectin-8 | | Authors: | Adrover Forteza, J, Puric, E, Nilsson, U.J, Anderluh, M, Logan, D.T. | | Deposit date: | 2024-07-03 | | Release date: | 2025-03-12 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Nanomolar inhibitor of the galectin-8 N-terminal domain binds via a non-canonical cation-pi interaction.
Commun Chem, 8, 2025
|
|
9FZF
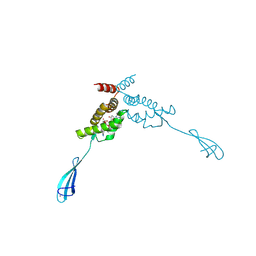 
 | | Transcriptional repressor NrdR from E. coli, ADP/dATP bound state | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Bimai, O, Rozman Grinberg, I, Sjoberg, B.M, Logan, D.T. | | Deposit date: | 2024-07-05 | | Release date: | 2025-03-19 | | Last modified: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (2.444 Å) | | Cite: | Bacterial transcriptional repressor NrdR - a flexible multifactorial nucleotide sensor.
Febs J., 292, 2025
|
|
6EOG
 
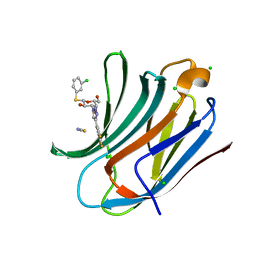 | | Human galectin-3c in complex with a galactose derivative | | Descriptor: | (2~{S},3~{R},4~{S},5~{R},6~{R})-2-(3-chlorophenyl)sulfanyl-6-(hydroxymethyl)-4-[4-[3,4,5-tris(fluoranyl)phenyl]-1,2,3-triazol-1-yl]oxane-3,5-diol, CHLORIDE ION, Galectin-3, ... | | Authors: | Hakansson, M, Nilsson, U.J, Zetterberg, F, Logan, D.T. | | Deposit date: | 2017-10-09 | | Release date: | 2018-08-22 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Monosaccharide Derivatives with Low-Nanomolar Lectin Affinity and High Selectivity Based on Combined Fluorine-Amide, Phenyl-Arginine, Sulfur-pi , and Halogen Bond Interactions.
ChemMedChem, 13, 2018
|
|
2J5A
 
 | | Folding of S6 structures with divergent amino-acid composition: pathway flexibility within partly overlapping foldons | | Descriptor: | 30S RIBOSOMAL PROTEIN S6, SODIUM ION | | Authors: | Hansson, S, Olofsson, L, Hedberg, L, Oliveberg, M, Logan, D.T. | | Deposit date: | 2006-09-13 | | Release date: | 2006-10-25 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Folding of S6 Structures with Divergent Amino Acid Composition: Pathway Flexibility within Partly Overlapping Foldons.
J.Mol.Biol., 365, 2007
|
|
2Y6J
 
 | | X-2 engineered mutated CBM4-2 Carbohydrate Binding Module from a Thermostable Rhodothermus marinus Xylanase | | Descriptor: | CALCIUM ION, XYLANASE | | Authors: | von Schantz, L, Hakansson, M, Logan, D.T, Walse, B, Osterlin, J, Nordberg-Karlsson, E, Ohlin, M. | | Deposit date: | 2011-01-24 | | Release date: | 2012-03-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for carbohydrate-binding specificity--a comparative assessment of two engineered carbohydrate-binding modules.
Glycobiology, 22, 2012
|
|
2Y6H
 
 | | X-2 L110F CBM4-2 Carbohydrate Binding Module from a Thermostable Rhodothermus marinus Xylanase | | Descriptor: | CALCIUM ION, XYLANASE | | Authors: | von Schantz, L, Hakansson, M, Logan, D.T, Walse, B, Osterlin, J, Nordberg-Karlsson, E, Ohlin, M. | | Deposit date: | 2011-01-21 | | Release date: | 2012-03-07 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Structural basis for carbohydrate-binding specificity--a comparative assessment of two engineered carbohydrate-binding modules.
Glycobiology, 22, 2012
|
|
3HFF
 
 | | Monomeric human Cu,Zn Superoxide dismutase without Zn ligands | | Descriptor: | Superoxide dismutase [Cu-Zn], ZINC ION | | Authors: | Saraboji, K, Nordlund, A, Leinartait, L, Oliveberg, M, Logan, D.T. | | Deposit date: | 2009-05-11 | | Release date: | 2009-06-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Functional features cause misfolding of the ALS-provoking enzyme SOD1.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
7BFG
 
 | |
7BFC
 
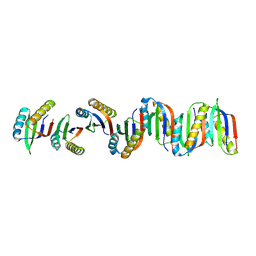 | |
7B90
 
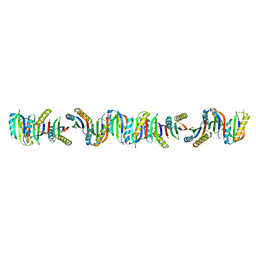 | | Circular permutant of ribosomal protein S6, P54-55 truncated, I8A mutant | | Descriptor: | 30S ribosomal protein S6,30S ribosomal protein S6 | | Authors: | Wang, H, Logan, D.T, Oliveberg, M. | | Deposit date: | 2020-12-13 | | Release date: | 2022-06-08 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Circular permutant of ribosomal protein S6, P54-55 truncate, I8A
To Be Published
|
|
7BFE
 
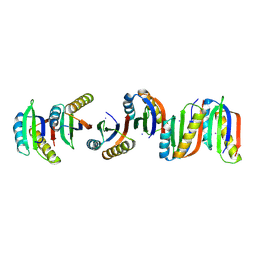 | |
7BFF
 
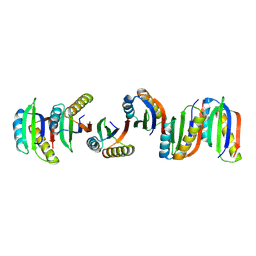 | |
7BFD
 
 | |
6Y4C
 
 | | Structure of galectin-3C in complex with lactose determined by serial crystallography using an XtalTool support | | Descriptor: | CHLORIDE ION, Galectin-3, beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Shilova, A, Hakansson, M, Welin, M, Kovacic, R, Mueller, U, Logan, D.T. | | Deposit date: | 2020-02-20 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Current status and future opportunities for serial crystallography at MAX IV Laboratory.
J.Synchrotron Radiat., 27, 2020
|
|
6Y78
 
 | | Structure of galectin-3C in complex with lactose determined by serial crystallography using a silicon nitride membrane support | | Descriptor: | Galectin-3, beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Hakansson, M, Welin, M, Shilova, A, Kovacic, R, Mueller, U, Logan, D.T. | | Deposit date: | 2020-02-28 | | Release date: | 2020-07-29 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Current status and future opportunities for serial crystallography at MAX IV Laboratory.
J.Synchrotron Radiat., 27, 2020
|
|
7B8V
 
 | |
3O0N
 
 | | Thermotoga maritima Ribonucleotide Reductase, NrdJ, in complex with dTTP and Adenosylcobalamin | | Descriptor: | 5'-DEOXYADENOSINE, CHLORIDE ION, COBALAMIN, ... | | Authors: | Larsson, K.-M, Logan, D.T, Nordlund, P. | | Deposit date: | 2010-07-19 | | Release date: | 2010-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural Basis for Adenosylcobalamin Activation in AdoCbl-Dependent Ribonucleotide Reductases.
Acs Chem.Biol., 5, 2010
|
|
