5FNE
 
 | |
5ANH
 
 | |
7B00
 
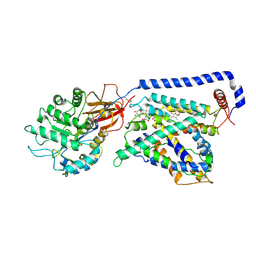 | | Human LAT2-4F2hc complex in the apo-state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Digitonin, Isoform 2 of 4F2 cell-surface antigen heavy chain, ... | | Authors: | Rodriguez, C.F, Escudero-Bravo, P, Garcia-Martin, C, Boskovic, J, Errasti-Murugarren, E, Palacin, M, Llorca, O. | | Deposit date: | 2020-11-17 | | Release date: | 2021-10-27 | | Last modified: | 2022-02-23 | | Method: | ELECTRON MICROSCOPY (3.98 Å) | | Cite: | Structural basis for substrate specificity of heteromeric transporters of neutral amino acids.
Proc Natl Acad Sci U S A, 118, 2021
|
|
7ZZK
 
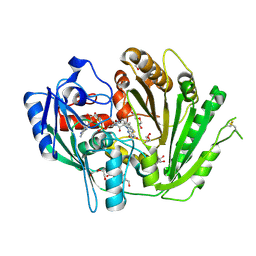 | | Structure of the N-acetyl-D-glucosamine oxidase from Ralstonia Solanacearum | | Descriptor: | ALANINE, DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Boverio, A, Rozeboom, H.J, Fraaije, M.W. | | Deposit date: | 2022-05-25 | | Release date: | 2022-08-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural Elucidation and Engineering of a Bacterial Carbohydrate Oxidase.
Biochemistry, 62, 2023
|
|
4R6W
 
 | |
4R6X
 
 | |
6FSK
 
 | |
6GBL
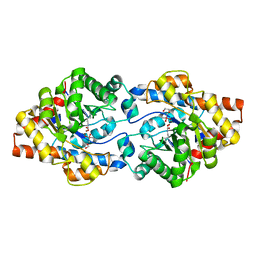 
 | | Repertoires of functionally diverse enzymes through computational design at epistatic active-site positions | | Descriptor: | 1,2-ETHANEDIOL, CACODYLATE ION, FORMIC ACID, ... | | Authors: | Khersonsky, O, Lipsh, R, Avizemer, Z, Goldsmith, M, Ashani, Y, Leader, H, Dym, O, Rogotner, S, Trudeau, D, Tawfik, D.S, Fleishman, S.J. | | Deposit date: | 2018-04-15 | | Release date: | 2018-10-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Automated Design of Efficient and Functionally Diverse Enzyme Repertoires.
Mol. Cell, 72, 2018
|
|
6GBJ
 
 | | Repertoires of functionally diverse enzymes through computational design at epistatic active-site positions | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, Parathion hydrolase, ... | | Authors: | Khersonsky, O, Lipsh, R, Avizemer, Z, Goldsmith, M, Ashani, Y, Leader, H, Dym, O, Rogotner, S, Trudeau, D, Tawfik, D.S, Fleishman, S.J. | | Deposit date: | 2018-04-15 | | Release date: | 2018-10-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Automated Design of Efficient and Functionally Diverse Enzyme Repertoires.
Mol. Cell, 72, 2018
|
|
6GBK
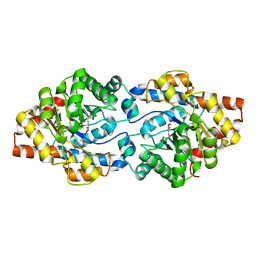 
 | | Repertoires of functionally diverse enzymes through computational design at epistatic active-site positions | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, FORMIC ACID, Parathion hydrolase, ... | | Authors: | Khersonsky, O, Lipsh, R, Avizemer, Z, Goldsmith, M, Ashani, Y, Leader, H, Dym, O, Rogotner, S, Trudeau, D, Tawfik, D.S, Fleishman, S.J. | | Deposit date: | 2018-04-15 | | Release date: | 2018-10-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Automated Design of Efficient and Functionally Diverse Enzyme Repertoires.
Mol. Cell, 72, 2018
|
|
4CZQ
 
 | |
4CZN
 
 | |
4CZR
 
 | |
4CZO
 
 | |
4CZP
 
 | |
