3QNT
 
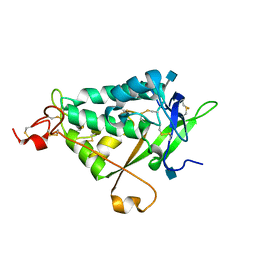 | | NPC1L1 (NTD) Structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Niemann-Pick C1-like protein 1 | | Authors: | Kwon, H.J. | | Deposit date: | 2011-02-09 | | Release date: | 2011-05-11 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | The Structure of the NPC1L1 N-Terminal Domain in a Closed Conformation.
Plos One, 6, 2011
|
|
3PDF
 
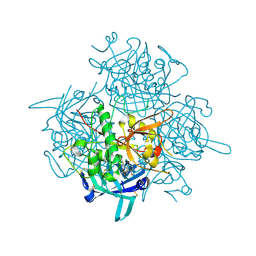 | | Discovery of Novel Cyanamide-Based Inhibitors of Cathepsin C | | Descriptor: | 2,5-dibromo-N-{(3R,5S)-1-[(Z)-iminomethyl]-5-methylpyrrolidin-3-yl}benzenesulfonamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Zhao, B, Laine, D. | | Deposit date: | 2010-10-22 | | Release date: | 2011-10-26 | | Last modified: | 2023-07-26 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Discovery of novel cyanamide-based inhibitors of cathepsin C.
Acs Med.Chem.Lett., 2, 2011
|
|
2M3K
 
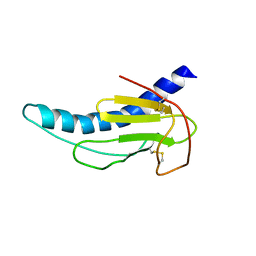 | |
2O5J
 
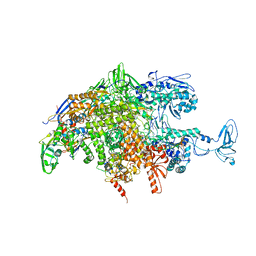 | |
2XQW
 
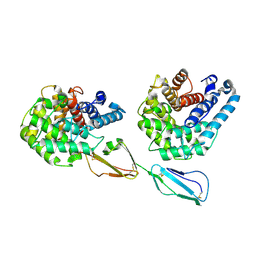 | | Structure of Factor H domains 19-20 in complex with complement C3d | | Descriptor: | COMPLEMENT C3, COMPLEMENT FACTOR H | | Authors: | Kajander, T, Lehtinen, M.J, Hyvarinen, S, Bhattacharjee, A, Leung, E, Isenman, D.E, Meri, S, Jokiranta, T.S, Goldman, A. | | Deposit date: | 2010-09-07 | | Release date: | 2011-02-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.306 Å) | | Cite: | Dual Interaction of Factor H with C3D and Glycosaminoglycans in Host-Nonhost Discrimination by Complement.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
5H6I
 
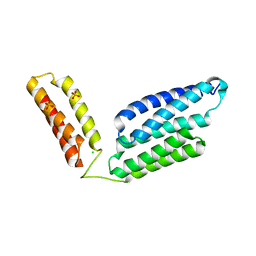 | | Crystal Structure of GBS CAMP Factor | | Descriptor: | CHLORIDE ION, Protein B, SULFATE ION | | Authors: | Jin, T.C, Brefo-Mensah, E.K. | | Deposit date: | 2016-11-13 | | Release date: | 2017-11-22 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal structure of theStreptococcus agalactiaeCAMP factor provides insights into its membrane-permeabilizing activity.
J.Biol.Chem., 293, 2018
|
|
3ZOJ
 
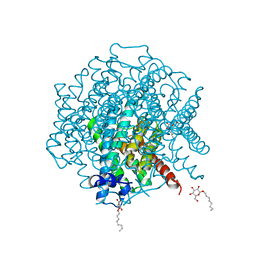 | | High-resolution structure of Pichia Pastoris aquaporin Aqy1 at 0.88 A | | Descriptor: | AQUAPORIN, CHLORIDE ION, octyl beta-D-glucopyranoside | | Authors: | Kosinska-Eriksson, U, Fischer, G, Friemann, R, Enkavi, G, Tajkhorshid, E, Neutze, R. | | Deposit date: | 2013-02-21 | | Release date: | 2013-06-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (0.88 Å) | | Cite: | Subangstrom Resolution X-Ray Structure Details Aquaporin-Water Interactions
Science, 340, 2013
|
|
