8JFS
 
 | | Phosphate bound acylphosphatase from Deinococcus radiodurans at 1 Angstrom resolution | | Descriptor: | 1,2-ETHANEDIOL, Acylphosphatase, CITRIC ACID, ... | | Authors: | Khakerwala, Z, Kumar, A, Makde, R.D. | | Deposit date: | 2023-05-18 | | Release date: | 2023-06-14 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Crystal structure of phosphate bound Acyl phosphatase mini-enzyme from Deinococcus radiodurans at 1 angstrom resolution.
Biochem.Biophys.Res.Commun., 671, 2023
|
|
8BV9
 
 | | Acylphosphatase from E. coli | | Descriptor: | 1,2-ETHANEDIOL, Acylphosphatase, GLYCEROL, ... | | Authors: | Gavira, J.A, Martinez-Rodriguez, S. | | Deposit date: | 2023-01-18 | | Release date: | 2023-10-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | First 3-D structural evidence of a native-like intertwined dimer in the acylphosphatase family.
Biochem.Biophys.Res.Commun., 682, 2023
|
|
7CHX
 
 | |
6KRB
 
 | | High resolution crystal structure of an Acylphosphatase protein cage | | Descriptor: | Acylphosphatase, GLYCEROL, SULFATE ION | | Authors: | Chatterjee, S, Nath, S, Sen, U. | | Deposit date: | 2019-08-21 | | Release date: | 2020-05-13 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.375 Å) | | Cite: | High resolution structure of Vibrio cholerae acylphosphatase (VcAcP) cage: Identification of drugs, location of its binding site and engineering to facilitate cage formation.
Biochem.Biophys.Res.Commun., 523, 2020
|
|
6CBU
 
 | | Crystal structure of C4S3: A computationally designed immunogen to target Carbohydrate-Occluded Epitopes on the HIV envelope | | Descriptor: | Acylphosphatase-1, SULFATE ION | | Authors: | Zhu, C, Ke, H.M, Swanstrom, R, Dokholyan, N.V. | | Deposit date: | 2018-02-05 | | Release date: | 2019-02-06 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Rationally designed carbohydrate-occluded epitopes elicit HIV-1 Env-specific antibodies.
Nat Commun, 10, 2019
|
|
4OJH
 | |
4OJG
 | |
4OJ3
 | |
4OJ1
 
 | |
4OIX
 
 | |
4HI2
 
 | |
4HI1
 
 | |
4G9I
 
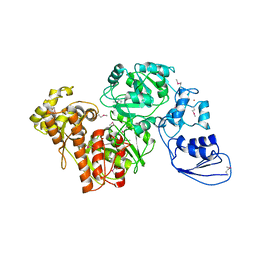 | | Crystal structure of T.kodakarensis HypF | | Descriptor: | Hydrogenase maturation protein HypF, ZINC ION | | Authors: | Tominaga, T, Watanabe, S, Matsumi, R, Atomi, H, Imanaka, T, Miki, K. | | Deposit date: | 2012-07-24 | | Release date: | 2012-10-24 | | Method: | X-RAY DIFFRACTION (4.5 Å) | | Cite: | Structure of the [NiFe]-hydrogenase maturation protein HypF from Thermococcus kodakarensis KOD1.
Acta Crystallogr.,Sect.F, 68, 2012
|
|
3VTI
 
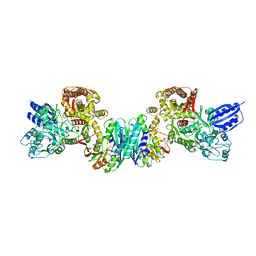 | | Crystal structure of HypE-HypF complex | | Descriptor: | FE (III) ION, Hydrogenase maturation factor, MAGNESIUM ION, ... | | Authors: | Shomura, Y, Higuchi, Y. | | Deposit date: | 2012-05-30 | | Release date: | 2012-07-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Structural basis for the reaction mechanism of S-carbamoylation of HypE by HypF in the maturation of [NiFe]-hydrogenases
J.Biol.Chem., 287, 2012
|
|
3VTH
 
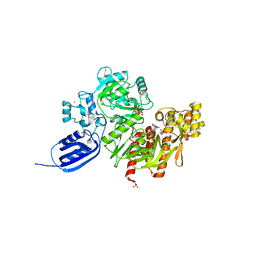 | |
3TRG
 
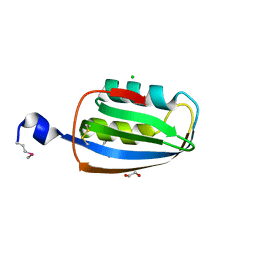 | | Structure of an acylphosphatase from Coxiella burnetii | | Descriptor: | 1,2-ETHANEDIOL, Acylphosphatase, CHLORIDE ION | | Authors: | Cheung, J, Franklin, M.C, Rudolph, M, Cassidy, M, Gary, E, Burshteyn, F, Love, J. | | Deposit date: | 2011-09-09 | | Release date: | 2011-10-19 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | Structural genomics for drug design against the pathogen Coxiella burnetii.
Proteins, 83, 2015
|
|
3TOQ
 
 | |
3TNV
 | |
3BR8
 | |
2W4P
 | | Human common-type acylphosphatase variant, A99G | | Descriptor: | ACYLPHOSPHATASE-1, GLYCEROL | | Authors: | Lam, S.Y, Sze, K.H, Wong, K.B. | | Deposit date: | 2008-11-29 | | Release date: | 2009-12-29 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A Rigidifying Salt-Bridge Favors the Activity of Thermophilic Enzyme at High Temperatures at the Expense of Low-Temperature Activity.
Plos Biol., 9, 2011
|
|
2W4D
 
 | | Acylphosphatase variant G91A from Pyrococcus horikoshii | | Descriptor: | ACYLPHOSPHATASE, PHOSPHATE ION, POTASSIUM ION | | Authors: | Lam, S.Y, Wong, K.B. | | Deposit date: | 2008-11-25 | | Release date: | 2009-12-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | A Rigidifying Salt-Bridge Favors the Activity of Thermophilic Enzyme at High Temperatures at the Expense of Low-Temperature Activity.
Plos Biol., 9, 2011
|
|
2W4C
 | | Human common-type acylphosphatase variant, A99 | | Descriptor: | ACYLPHOSPHATASE-1 | | Authors: | Lam, S.Y, Wong, K.B. | | Deposit date: | 2008-11-25 | | Release date: | 2009-12-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | A Rigidifying Salt-Bridge Favors the Activity of Thermophilic Enzyme at High Temperatures at the Expense of Low-Temperature Activity.
Plos Biol., 9, 2011
|
|
2VH7
 | |
2LXF
 | |
2K7K
 
 | | Human Acylphosphatase (AcPh) common type | | Descriptor: | Acylphosphatase-1 | | Authors: | Gribenko, A.V, Patel, M.M, Liu, J, McCallum, S.A, Wang, C, Makhatadze, G.I. | | Deposit date: | 2008-08-13 | | Release date: | 2009-02-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Rational stabilization of enzymes by computational redesign of surface charge-charge interactions
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
