9L2P
 
 | |
9L34
 
 | | Crystal structure of AnAE-H270A mutant | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Cellulose-binding GDSL lipase/acylhydrolase, ... | | Authors: | Xing, S.Q, Hu, G.L, He, L.P. | | Deposit date: | 2024-12-17 | | Release date: | 2025-10-29 | | Last modified: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Structural and functional characterization of a novel GDSL-acetylesterase from Aspergillus niger reveals unique acylated choline specificity
Chem Eng J, 530, 2026
|
|
9L37
 
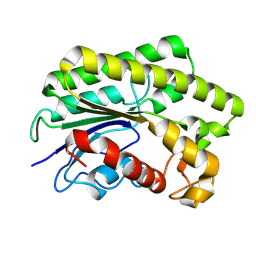 | | Crystal structure of AnAE-N267D-A101F mutant | | Descriptor: | Cellulose-binding GDSL lipase/acylhydrolase | | Authors: | Xing, S.Q, Hu, G.L, He, L.P. | | Deposit date: | 2024-12-17 | | Release date: | 2025-10-29 | | Last modified: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | Structural and functional characterization of a novel GDSL-acetylesterase from Aspergillus niger reveals unique acylated choline specificity
Chem Eng J, 530, 2026
|
|
9L39
 
 | | Crystal structure of AnAE-N267D-I139D mutant | | Descriptor: | Cellulose-binding GDSL lipase/acylhydrolase | | Authors: | Xing, S.Q, Hu, G.L, He, L.P. | | Deposit date: | 2024-12-17 | | Release date: | 2025-10-29 | | Last modified: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Structural and functional characterization of a novel GDSL-acetylesterase from Aspergillus niger reveals unique acylated choline specificity
Chem Eng J, 530, 2026
|
|
9L3A
 
 | | Crystal structure of AnAE-N267D-Y102W mutant | | Descriptor: | Cellulose-binding GDSL lipase/acylhydrolase | | Authors: | Xing, S.Q, Hu, G.L, He, L.P. | | Deposit date: | 2024-12-17 | | Release date: | 2025-10-29 | | Last modified: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Structural and functional characterization of a novel GDSL-acetylesterase from Aspergillus niger reveals unique acylated choline specificity
Chem Eng J, 530, 2026
|
|
9L3B
 
 | | Crystal structure of AnAE-N267D-Y269W mutant | | Descriptor: | Cellulose-binding GDSL lipase/acylhydrolase | | Authors: | Xing, S.Q, Hu, G.L, He, L.P. | | Deposit date: | 2024-12-18 | | Release date: | 2025-10-29 | | Last modified: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Structural and functional characterization of a novel GDSL-acetylesterase from Aspergillus niger reveals unique acylated choline specificity
Chem Eng J, 530, 2026
|
|
9L62
 
 | |
9L63
 
 | |
9L64
 
 | |
9L65
 
 | |
9L7W
 
 | |
9V73
 
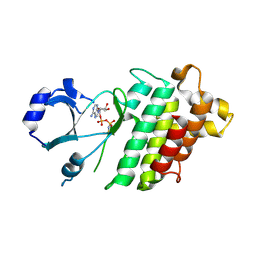 | |
9V79
 
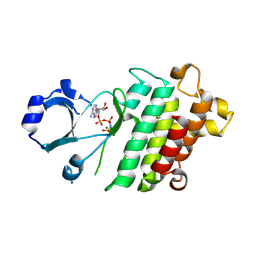 | |
9WLP
 
 | |
9WYQ
 
 | |
9WYR
 
 | |
9WYS
 
 | |
