8E22
 
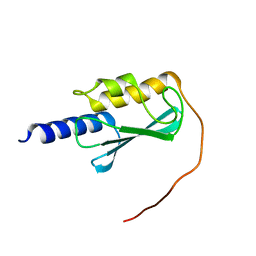 | | VPS37A_21-148 | | Descriptor: | Vacuolar protein sorting-associated protein 37A | | Authors: | Tian, F, Ye, Y.S. | | Deposit date: | 2022-08-12 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Identification of membrane curvature sensing motifs essential for VPS37A phagophore recruitment and autophagosome closure.
Commun Biol, 7, 2024
|
|
8E2O
 
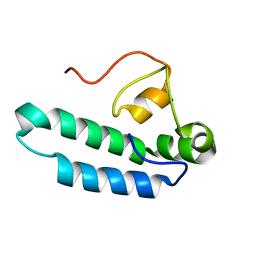 | |
8E4V
 
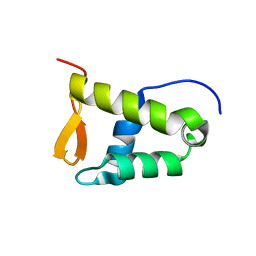 | | Solution structure of the WH domain of MORF | | Descriptor: | Isoform 3 of Histone acetyltransferase KAT6B | | Authors: | Zhang, Y, Kutateladze, T.G. | | Deposit date: | 2022-08-19 | | Release date: | 2023-05-10 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | MORF and MOZ acetyltransferases target unmethylated CpG islands through the winged helix domain.
Nat Commun, 14, 2023
|
|
8E6Y
 
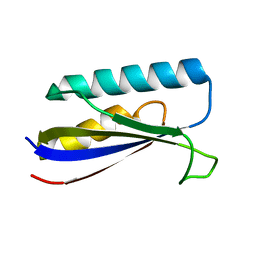 | |
8EB1
 
 | | Chim2 - Intragenic antimicrobial peptide | | Descriptor: | Unconventional myosin-Ih, Transcription activator BRG1 intragenic antimicrobial chimeric peptide | | Authors: | de Freitas, T.V, Oliveira, A.L, Santos, M.A, Brand, G.D. | | Deposit date: | 2022-08-30 | | Release date: | 2023-03-01 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Release of immunomodulatory peptides at bacterial membrane interfaces as a novel strategy to fight microorganisms.
J.Biol.Chem., 299, 2023
|
|
8EF4
 
 | |
8ENP
 
 | | UBE3A isoform 3 AZUL | | Descriptor: | Isoform III of Ubiquitin-protein ligase E3A, ZINC ION | | Authors: | Bregnard, T.A, Bezsonova, I. | | Deposit date: | 2022-09-30 | | Release date: | 2023-10-18 | | Last modified: | 2025-02-05 | | Method: | SOLUTION NMR | | Cite: | Differences in structure, dynamics, and zinc coordination between isoforms of human ubiquitin ligase UBE3A.
J.Biol.Chem., 301, 2024
|
|
8EO9
 
 | | The solution structure of abxF, an enzyme catalyzing the formation of chiral spiroketal of an antibiotics, (-)-ABX | | Descriptor: | Glyoxalase | | Authors: | Jia, X, Yan, X, Mobli, M, Qu, X. | | Deposit date: | 2022-10-02 | | Release date: | 2024-04-03 | | Last modified: | 2025-04-16 | | Method: | SOLUTION NMR | | Cite: | The solution structure of abxF, an enzyme catalyzing the formation of chiral spiroketal of an antibiotics, (-)-ABX.
Nat.Chem., 2025
|
|
8EOD
 
 | |
8EP5
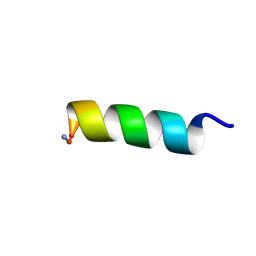 
 | | Solution NMR structure of a computationally designed mastoparan-like peptide, mastoparan-R1 | | Descriptor: | Mastoparan-R1 peptide | | Authors: | Freitas, C.D.P, Oshiro, K.G.N, Macedo, M.L.R, Cardoso, M.H, Franco, O.L, Liao, L.M. | | Deposit date: | 2022-10-05 | | Release date: | 2023-10-18 | | Last modified: | 2024-11-13 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of a computationally designed mastoparan-like peptide, mastoparan-R1.
To Be Published
|
|
8EPT
 
 | | UBE3A isoform 2 AZUL domain | | Descriptor: | Ubiquitin-protein ligase E3A, ZINC ION | | Authors: | Bregnard, T.A, Bezsonova, I. | | Deposit date: | 2022-10-06 | | Release date: | 2023-07-05 | | Last modified: | 2025-02-05 | | Method: | SOLUTION NMR | | Cite: | Differences in structure, dynamics, and zinc coordination between isoforms of human ubiquitin ligase UBE3A.
J.Biol.Chem., 301, 2024
|
|
8EPY
 
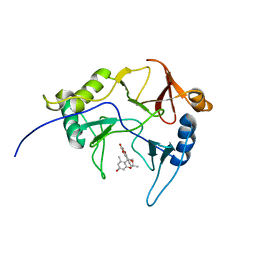 | | The solution structure of abxF in complex with its product (-)-ABX, an enzyme catalyzing the formation of the chiral spiroketal of an anthrabenzoxocinone antibiotic, (-)-ABX | | Descriptor: | (6R,16R)-3,11,13,15-tetrahydroxy-1,6,9,9-tetramethyl-6,7,9,16-tetrahydro-14H-6,16-epoxyanthra[2,3-e]benzo[b]oxocin-14-one, Glyoxalase | | Authors: | Jia, X, Yan, X, Qu, X, Mobli, M. | | Deposit date: | 2022-10-06 | | Release date: | 2024-04-10 | | Last modified: | 2025-04-23 | | Method: | SOLUTION NMR | | Cite: | An enzymatic dual-oxa Diels–Alder reaction constructs the oxygen-bridged tricyclic acetal unit of (–)-anthrabenzoxocinone
Nat.Chem., 2025
|
|
8ERU
 | | Solution NMR structure of a computationally designed mastoparan-like peptide, mastoparan-R4 | | Descriptor: | Mastoparan-R4 peptide | | Authors: | Freitas, C.D.P, Oshiro, K.G.N, Macedo, M.L.R, Cardoso, M.H, Franco, O.L, Liao, L.M. | | Deposit date: | 2022-10-12 | | Release date: | 2023-10-18 | | Last modified: | 2024-11-20 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of a computacionally designed mastoparan-like peptide, mastoparan-R4
To Be Published
|
|
8ERY
 
 | |
8ERZ
 
 | |
8ES0
 
 | |
8ES1
 | |
8ES2
 | |
8ES3
 
 | |
8F04
 
 | |
8F2F
 | |
8F36
 
 | |
8F37
 
 | |
8F4V
 
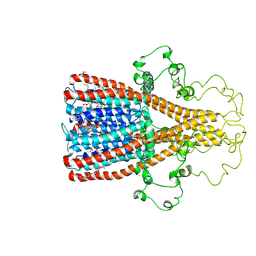 | | Alpha7 nicotinic acetylcholine receptor intracellular and transmembrane domains bound to ivermectin in a desensitized state | | Descriptor: | (2aE,4E,5'S,6S,6'R,7S,8E,11R,13R,15S,17aR,20R,20aR,20bS)-6'-[(2S)-butan-2-yl]-20,20b-dihydroxy-5',6,8,19-tetramethyl-17 -oxo-3',4',5',6,6',10,11,14,15,17,17a,20,20a,20b-tetradecahydro-2H,7H-spiro[11,15-methanofuro[4,3,2-pq][2,6]benzodioxacy clooctadecine-13,2'-pyran]-7-yl 2,6-dideoxy-4-O-(2,6-dideoxy-3-O-methyl-alpha-L-arabino-hexopyranosyl)-3-O-methyl-alpha-L-arabino-hexopyranoside, Neuronal acetylcholine receptor subunit alpha-7 | | Authors: | Bondarenko, V, Chen, Q, Tang, P. | | Deposit date: | 2022-11-11 | | Release date: | 2023-02-22 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural Elucidation of Ivermectin Binding to alpha 7nAChR and the Induced Channel Desensitization.
Acs Chem Neurosci, 14, 2023
|
|
8FA3
 
 | |
