2LPC
 
 | | NMR STRUCTURE of Bcl-XL | | Descriptor: | Bcl-2-like protein 1 | | Authors: | Wysoczanski, P, Mart, R.J, Loveridge, J.E, Williams, C, Whittaker, S.B.-M, Crump, M.P, Allemann, R.K. | | Deposit date: | 2012-02-09 | | Release date: | 2012-03-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of a Photoswitchable Apoptosis Activating Bak Peptide Bound to Bcl-x(L).
J.Am.Chem.Soc., 134, 2012
|
|
2LPD
 
 | |
2LPE
 
 | |
2LPF
 
 | | R state structure of monomeric phospholamban (C36A, C41F, C46A) | | Descriptor: | Cardiac phospholamban | | Authors: | De Simone, A, Montalvao, R.W, Gustavsson, M, Shi, L, Veglia, G, Vendruscolo, M. | | Deposit date: | 2012-02-11 | | Release date: | 2013-02-13 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structures of the Excited States of Phospholamban and Shifts in Their Populations upon Phosphorylation.
Biochemistry, 52, 2013
|
|
2LPI
 | | NMR structure of a monomeric mutant (A72R) of major ampullate spidroin 1 N-terminal domain | | Descriptor: | Major ampullate spidroin 1 | | Authors: | Jaudzems, K, Nordling, K, Landreh, M, Rising, A, Askarieh, G, Knight, S.D, Johansson, J. | | Deposit date: | 2012-02-14 | | Release date: | 2012-06-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | pH-Dependent Dimerization of Spider Silk N-Terminal Domain Requires Relocation of a Wedged Tryptophan Side Chain.
J.Mol.Biol., 422, 2012
|
|
2LPJ
 
 | | NMR structure of major ampullate spidroin 1 N-terminal domain at pH 7.2 | | Descriptor: | Major ampullate spidroin 1 | | Authors: | Jaudzems, K, Nordling, K, Landreh, M, Rising, A, Askarieh, G, Knight, S.D, Johansson, J. | | Deposit date: | 2012-02-14 | | Release date: | 2012-06-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | pH-Dependent Dimerization of Spider Silk N-Terminal Domain Requires Relocation of a Wedged Tryptophan Side Chain.
J.Mol.Biol., 422, 2012
|
|
2LPK
 
 | | Solution NMR of the specialized apo-acyl carrier protein (RPA2022) from Rhodopseudomonas palustris, Northeast Structural Genomics Consortium Target RpR324 | | Descriptor: | Specialized acyl carrier protein | | Authors: | Ramelot, T.A, Rossi, P, Yang, Y, Lee, H, Ertekin, A, Wang, H, Ciccosanti, C, Kohan, E, Maglaqui, M, Janjua, H, Nair, R, Rost, B, Acton, T.B, Xiao, R, Everett, J.K, Prestegard, J.H, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2012-02-14 | | Release date: | 2012-02-29 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure of a specialized acyl carrier protein essential for lipid A biosynthesis with very long-chain fatty acids in open and closed conformations.
Biochemistry, 51, 2012
|
|
2LPM
 
 | |
2LPN
 
 | | Solution Structure of N-Terminal domain of human Conserved Dopamine Neurotrophic Factor (CDNF) | | Descriptor: | Cerebral dopamine neurotrophic factor | | Authors: | Latge, C, Cabral, K.M.S, Foguel, D, Pires, J.R.M, Almeida, M.S. | | Deposit date: | 2012-02-15 | | Release date: | 2013-02-20 | | Last modified: | 2013-05-22 | | Method: | SOLUTION NMR | | Cite: | (1)H-, (13)C- and (15)N-NMR assignment of the N-terminal domain of human cerebral dopamine neurotrophic factor (CDNF).
Biomol.Nmr Assign., 7, 2013
|
|
2LPS
 
 | |
2LPT
 
 | |
2LPU
 | | Solution structures of KmAtg10 | | Descriptor: | KmAtg10 | | Authors: | Yamaguchi, M, Noda, N.N, Yamamoto, H, Shima, T, Kumeta, H, Kobashigawa, Y, Akada, R, Ohsumi, Y, Inagaki, F. | | Deposit date: | 2012-02-19 | | Release date: | 2012-08-01 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural insights into atg10-mediated formation of the autophagy-essential atg12-atg5 conjugate
Structure, 20, 2012
|
|
2LPV
 
 | |
2LPW
 
 | | human CEB25 minisatellite G-quadruplex | | Descriptor: | DNA (26-MER) | | Authors: | Amrane, S, Adrian, M, Heddi, B, Serero, A, Nicolas, A, Mergny, J.L, Phan, A.T. | | Deposit date: | 2012-02-20 | | Release date: | 2012-03-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Formation of Pearl-Necklace Monomorphic G-Quadruplexes in the Human CEB25 Minisatellite.
J.Am.Chem.Soc., 134, 2012
|
|
2LPX
 
 | | Solution Structure of Strawberry Allergen Fra a 1e | | Descriptor: | Major strawberry allergen Fra a 1-E | | Authors: | Seutter von Loetzen, C, Hartl-Spiegelhauer, O, Schweimer, K, Schwab, W, Roesch, P. | | Deposit date: | 2012-02-20 | | Release date: | 2012-03-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the strawberry allergen Fra a 1.
Biosci.Rep., 32, 2012
|
|
2LPZ
 | | Atomic model of the Type-III Secretion System Needle | | Descriptor: | Protein prgI | | Authors: | Loquet, A, Sgourakis, N.G, Gupta, R, Giller, K, Riedel, D, Goosmann, C, Griesinger, C, Kolbe, M.G, Baker, D, Becker, S, Lange, A. | | Deposit date: | 2012-02-21 | | Release date: | 2012-05-16 | | Last modified: | 2024-05-15 | | Method: | SOLID-STATE NMR | | Cite: | Atomic model of the type III secretion system needle.
Nature, 486, 2012
|
|
2LQ0
 
 | | Solution structure of de novo designed antifreeze peptide 1m | | Descriptor: | de novo designed antifreeze peptide 1m | | Authors: | Bhunia, A. | | Deposit date: | 2012-02-21 | | Release date: | 2012-10-24 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structures, dynamics, and ice growth inhibitory activity of Peptide fragments derived from an antarctic yeast protein
Plos One, 7, 2012
|
|
2LQ1
 
 | | Solution structure of de novo designed antifreeze peptide 3 | | Descriptor: | de novo designed antifreeze peptide 3 | | Authors: | Bhunia, A. | | Deposit date: | 2012-02-21 | | Release date: | 2012-10-24 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structures, dynamics, and ice growth inhibitory activity of Peptide fragments derived from an antarctic yeast protein
Plos One, 7, 2012
|
|
2LQ2
 
 | | Solution structure of de novo designed peptide 4m | | Descriptor: | de novo designed antifreeze peptide 4m | | Authors: | Bhunia, A. | | Deposit date: | 2012-02-22 | | Release date: | 2012-10-24 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structures, dynamics, and ice growth inhibitory activity of Peptide fragments derived from an antarctic yeast protein
Plos One, 7, 2012
|
|
2LQ3
 
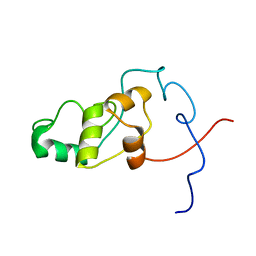 | | Solution NMR Structure of syc0711_d from Synechococcus sp., Northeast Structural Genomics Consortium (NESG) Target SnR212 | | Descriptor: | Uncharacterized protein | | Authors: | Mills, J.L, Sathyamoorthy, B, Pulavarti, S, Janjua, H, Kohan, E, Wang, D, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2012-02-23 | | Release date: | 2012-03-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of syc0711_d from Synechococcus sp., Northeast Structural Genomics Consortium (NESG) Target SnR212
To be Published
|
|
2LQ4
 
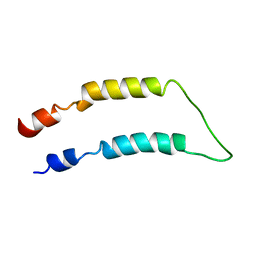 | |
2LQ5
 
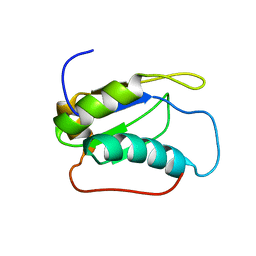 | |
2LQ6
 
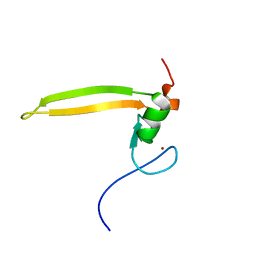 | | Solution structure of BRD1 PHD2 finger | | Descriptor: | Bromodomain-containing protein 1, ZINC ION | | Authors: | Liu, L, Wu, J. | | Deposit date: | 2012-02-25 | | Release date: | 2012-10-24 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of an atypical PHD finger in BRPF2 and its interaction with DNA
J.Struct.Biol., 180, 2012
|
|
2LQ7
 
 | |
2LQ8
 
 | | Domain interaction in Thermotoga maritima NusG | | Descriptor: | Transcription antitermination protein nusG | | Authors: | Droegemueller, J, Stegmann, C, Burmann, B, Roesch, P, Wahl, M.C, Schweimer, K. | | Deposit date: | 2012-02-27 | | Release date: | 2013-01-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | An Autoinhibited State in the Structure of Thermotoga maritima NusG.
Structure, 21, 2013
|
|
