1QWB
 
 | |
1QWV
 
 | |
1QX9
 
 | |
1QXB
 
 | | NMR structure determination of the self complementary DNA Dodecamer CGCGAATT*CGCG in which a ribose is inserted between the 3'-OH of T8 and the 5'-phosphate group of C9 | | Descriptor: | 5'-d(CpGpCpGpApApTpTpCpGpCpG)-3', beta-D-ribofuranose | | Authors: | Nauwelaerts, K, Vastmans, K, Froeyen, M, Kempeneers, V, Rozenski, J, Rosemeyer, H, Van Aerschot, A, Busson, R, Efimtseva, E, Mikhailov, S, Lescrinier, E, Herdewijn, P. | | Deposit date: | 2003-09-05 | | Release date: | 2004-02-03 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Cleavage of DNA without loss of genetic information by incorporation of a disaccharide nucleoside.
Nucleic Acids Res., 31, 2003
|
|
1QXC
 
 | | NMR structure of the fragment 25-35 of beta amyloid peptide in 20/80 v:v hexafluoroisopropanol/water mixture | | Descriptor: | 11-mer peptide from Amyloid beta A4 protein | | Authors: | D'Ursi, A.M, Armenante, M.R, Guerrini, R, Salvadori, S, Sorrentino, G, Picone, D. | | Deposit date: | 2003-09-05 | | Release date: | 2004-09-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of amyloid beta-peptide (25-35) in different media
J.Med.Chem., 47, 2004
|
|
1QXF
 
 | | SOLUTION STRUCTURE OF 30S RIBOSOMAL PROTEIN S27E FROM ARCHAEOGLOBUS FULGIDUS: GR2, A NESG TARGET PROTEIN | | Descriptor: | 30S RIBOSOMAL PROTEIN S27E | | Authors: | Herve Du Penhoat, C, Atreya, H.S, Shen, Y, Liu, G, Acton, T.B, Xiao, R, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2003-09-05 | | Release date: | 2003-09-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The NMR solution structure of the 30S ribosomal protein S27e encoded in gene RS27_ARCFU of Archaeoglobus fulgidis reveals a novel protein fold
Protein Sci., 13, 2004
|
|
1QXN
 
 | | Solution Structure of the 30 kDa Polysulfide-sulfur Transferase Homodimer from Wolinella Succinogenes | | Descriptor: | PENTASULFIDE-SULFUR, sulfide dehydrogenase | | Authors: | Lin, Y.J, Dancea, F, Loehr, F, Klimmek, O, Pfeiffer-Marek, S, Nilges, M, Wienk, H, Kroeger, A, Rueterjans, H. | | Deposit date: | 2003-09-08 | | Release date: | 2004-02-24 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the 30 kDa Polysulfide-Sulfur Transferase Homodimer from Wolinella succinogenes
Biochemistry, 43, 2004
|
|
1QXQ
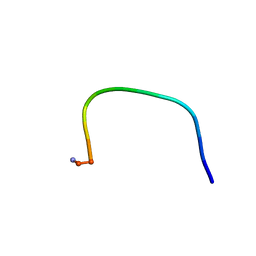 
 | |
1QYT
 
 | | Solution structure of fragment (25-35) of beta amyloid peptide in SDS micellar solution | | Descriptor: | 11-mer peptide from Amyloid beta A4 protein | | Authors: | D'Ursi, A.M, Armenante, M.R, Guerrini, R, Salvadori, S, Sorrentino, G, Picone, D. | | Deposit date: | 2003-09-12 | | Release date: | 2004-12-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of amyloid beta-peptide (25-35) in different media
J.Med.Chem., 47, 2004
|
|
1QZE
 
 | | HHR23a protein structure based on residual dipolar coupling data | | Descriptor: | UV excision repair protein RAD23 homolog A | | Authors: | Walters, K.J, Lech, P.J, Goh, A.M, Wang, Q, Howley, P.M. | | Deposit date: | 2003-09-16 | | Release date: | 2003-10-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | DNA-repair protein hHR23a alters its protein structure upon binding proteasomal subunit S5a
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1QZP
 
 | |
1R02
 
 | |
1R05
 
 | | Solution Structure of Max B-HLH-LZ | | Descriptor: | Max protein | | Authors: | Sauv, S, Tremblay, L, Lavigne, P. | | Deposit date: | 2003-09-19 | | Release date: | 2003-10-21 | | Last modified: | 2024-11-13 | | Method: | SOLUTION NMR | | Cite: | The NMR solution structure of a mutant of the Max b/HLH/LZ free of DNA: insights into the specific and reversible DNA binding mechanism of dimeric transcription factors
J.Mol.Biol., 342, 2004
|
|
1R1F
 
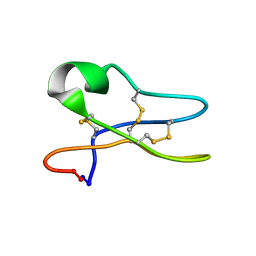 | | Solution Structure of the Cyclotide Palicourein: Implications for the development of pharmaceutical and agricultural applications | | Descriptor: | Palicourein | | Authors: | Barry, D.G, Daly, N.L, Bokesch, H.R, Gustafson, K.R, Craik, D.J. | | Deposit date: | 2003-09-23 | | Release date: | 2004-04-06 | | Last modified: | 2024-11-20 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the cyclotide palicourein: implications for the development of a pharmaceutical framework.
STRUCTURE, 12, 2004
|
|
1R21
 
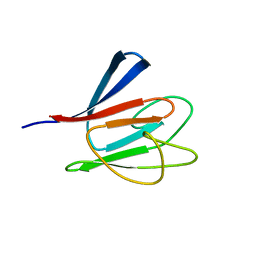 | | Solution Structure of human Ki67 FHA Domain | | Descriptor: | Antigen Ki-67 | | Authors: | Byeon, I.J, Li, H, Tsai, M.D. | | Deposit date: | 2003-09-25 | | Release date: | 2003-12-30 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of Human Ki67 FHA Domain and its Binding to a Phosphoprotein Fragment from hNIFK Reveal Unique Recognition Sites and New Views to the Structural Basis of FHA Domain Functions
J.Mol.Biol., 335, 2004
|
|
1R2A
 
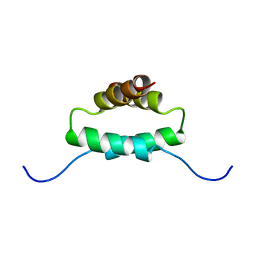 | | THE MOLECULAR BASIS FOR PROTEIN KINASE A ANCHORING REVEALED BY SOLUTION NMR | | Descriptor: | PROTEIN (CAMP-DEPENDENT PROTEIN KINASE TYPE II REGULATORY SUBUNIT) | | Authors: | Newlon, M.G, Roy, M, Morikis, D, Hausken, Z.E, Coghlan, V, Scott, J.D, Jennings, P.A. | | Deposit date: | 1998-12-07 | | Release date: | 1998-12-16 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The molecular basis for protein kinase A anchoring revealed by solution NMR.
Nat.Struct.Biol., 6, 1999
|
|
1R2L
 
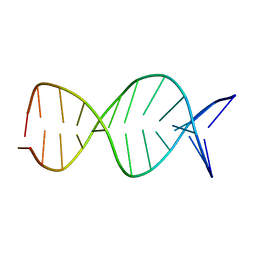 | | A parallel stranded DNA duplex with an A-G mismatch base-pair | | Descriptor: | 5'-D(P*CP*CP*TP*AP*TP*GP*AP*AP*AP*TP*CP*C)-3', DNA (5'-D(P*(DNR)P*(DNR)P*DAP*DTP*DAP*DAP*DTP*DTP*DTP*DAP*(DNR)P*(DNR))-3') | | Authors: | Venkitakrishnan, R.P, Bhaumik, S.R, Chary, K.V.R, govil, G, Liu, K, Howard, F.B, Miles, T.H. | | Deposit date: | 2003-09-29 | | Release date: | 2004-10-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A parallel stranded DNA duplex with an A-G mismatch base-pair: (CCATAATTTACC:CCTATGAAATCC)
RECENT TRENDS IN BIOPHYS.RES., 2004
|
|
1R2N
 
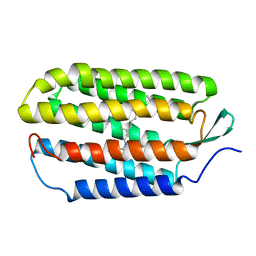 | | NMR structure of the all-trans retinal in dark-adapted Bacteriorhodopsin | | Descriptor: | Bacteriorhodopsin, RETINAL | | Authors: | Patzelt, H, Simon, B, terLaak, A, Kessler, B, Kuhne, R, Schmieder, P, Oesterhaelt, D, Oschkinat, H. | | Deposit date: | 2003-09-29 | | Release date: | 2003-10-28 | | Last modified: | 2024-11-13 | | Method: | SOLUTION NMR | | Cite: | The structures of the active center in dark-adapted bacteriorhodopsin by solution-state NMR spectroscopy
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
1R2P
 
 | | Solution structure of domain 5 from the ai5(gamma) group II intron | | Descriptor: | 34-MER | | Authors: | Sigel, R.K.O, Sashital, D.G, Abramovitz, D.L, Palmer III, A.G, Butcher, S.E, Pyle, A.M. | | Deposit date: | 2003-09-29 | | Release date: | 2004-02-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of domain 5 of a group II intron ribozyme reveals a new RNA motif.
Nat.Struct.Mol.Biol., 11, 2004
|
|
1R2U
 
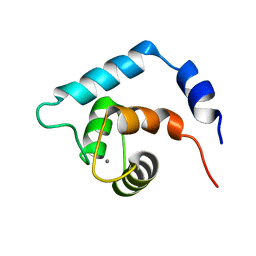 | | NMR structure of the N domain of trout cardiac troponin C at 30 C | | Descriptor: | CALCIUM ION, troponin C | | Authors: | Blumenschein, T.M, Gillis, T.E, Tibbits, G.F, Sykes, B.D. | | Deposit date: | 2003-09-29 | | Release date: | 2004-06-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Effect of temperature on the structure of trout troponin C
Biochemistry, 43, 2004
|
|
1R36
 
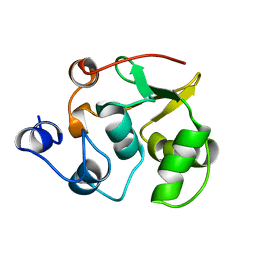 | | NMR-based structure of autoinhibited murine Ets-1 deltaN301 | | Descriptor: | C-ets-1 protein | | Authors: | Lee, G.M, Donaldson, L.W, Pufall, M.A, Kang, H.-S, Pot, I, Graves, B.J, McIntosh, L.P. | | Deposit date: | 2003-09-30 | | Release date: | 2004-11-09 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | The Structural and Dynamic Basis of Ets-1 DNA Binding Autoinhibition
J.Biol.Chem., 280, 2005
|
|
1R48
 
 | | Solution structure of the C-terminal cytoplasmic domain residues 468-497 of Escherichia coli protein ProP | | Descriptor: | Proline/betaine transporter | | Authors: | Zoetewey, D.L, Tripet, B.P, Kutateladze, T.G, Overduin, M.J, Wood, J.M, Hodges, R.S. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-23 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the C-terminal Antiparallel Coiled-coil Domain from Escherichia coli Osmosensor ProP.
J.Mol.Biol., 334, 2003
|
|
1R4D
 
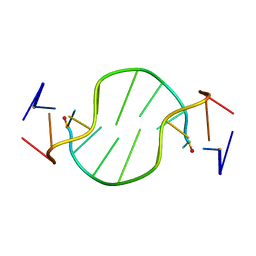 | | Solution structure of the chimeric L/D DNA oligonucleotide d(C8metGCGC(L)G(L)CGCG)2 | | Descriptor: | 5'-D(*CP*(8MG)P*CP*GP*(0DC)P*(0DG)P*CP*GP*CP*G)-3' | | Authors: | Cherrak, I, Mauffret, O, Santamaria, F, Rayner, B, Hocquet, A, Ghomi, M, Fermandjian, S. | | Deposit date: | 2003-10-06 | | Release date: | 2003-10-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | L-nucleotides and 8-methylguanine of d(C1m8G2C3G4C5LG6LC7G8C9G10)2 act cooperatively to promote a left-handed helix under physiological salt conditions.
Nucleic Acids Res., 31, 2003
|
|
1R4H
 
 | | NMR Solution structure of the IIIc domain of GB Virus B IRES Element | | Descriptor: | 5'-R(*GP*GP*GP*CP*AP*AP*GP*CP*CP*C)-3' | | Authors: | Kaluarachchi, K, Thiviyanathan, V, Rijinbrand, R, Lemon, S.M, Gorenstein, D.G. | | Deposit date: | 2003-10-06 | | Release date: | 2004-10-19 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Mutational and structural analysis of stem-loop IIIC of the hepatitis C virus and GB virus B internal ribosome entry sites.
J.Mol.Biol., 343, 2004
|
|
1R4K
 
 | | Solution Structure of the Drosophila Argonaute 1 PAZ Domain | | Descriptor: | Argonaute 1 | | Authors: | Yan, K.S, Yan, S, Farooq, A, Han, A, Zeng, L, Zhou, M.-M. | | Deposit date: | 2003-10-07 | | Release date: | 2003-12-09 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure and conserved RNA binding of the PAZ domain
Nature, 426, 2003
|
|
