10AD
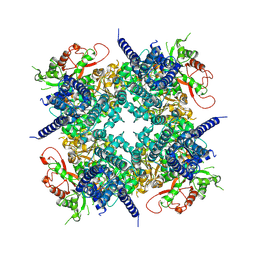 
 | | Cryo-EM structure of the human BK channel bound to the agonist NS1619 | | Descriptor: | 3-[2-oxidanyl-5-(trifluoromethyl)phenyl]-6-(trifluoromethyl)-1~{H}-benzimidazol-2-one, CALCIUM ION, Isoform 5 of Calcium-activated potassium channel subunit alpha-1, ... | | Authors: | Gonzalez-Sanabria, N, Contreras, G.F, Perozo, E, Latorre, R. | | Deposit date: | 2026-01-08 | | Release date: | 2026-02-04 | | Last modified: | 2026-02-11 | | Method: | ELECTRON MICROSCOPY (3.44 Å) | | Cite: | The BK channel-NS1619 agonist complex reveals molecular insights into allosteric activation gating.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
10AY
 
 | | Cryo-EM structure of CRBN-DDB1 in complex with HBS1L and TNG961 | | Descriptor: | DNA damage-binding protein 1, HBS1-like protein, N-{6-[(3R)-2,6-dioxopiperidin-3-yl]naphthalen-1-yl}-N'-{2-[6-(trifluoromethyl)-1-benzothiophen-2-yl]propan-2-yl}urea, ... | | Authors: | Whittington, D.A. | | Deposit date: | 2026-01-09 | | Release date: | 2026-04-22 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | TNG961 is a selective oral HBS1L molecular glue degrader for the treatment of FOCAD-deleted cancers
Cancer Discov, 2026
|
|
10FZ
 
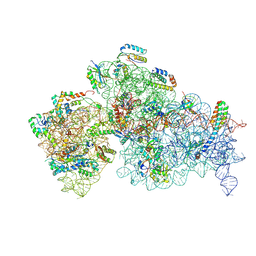 | | 30S ribosomal subunit from E. coli missing the gene encoding for the 16S rRNA 2'-O-methyltransferase RsmI | | Descriptor: | 16S rRNA, MAGNESIUM ION, Small ribosomal subunit protein bS16, ... | | Authors: | Barmada, M.I, Nandi, S, Conn, G.L. | | Deposit date: | 2026-01-18 | | Release date: | 2026-03-18 | | Last modified: | 2026-04-08 | | Method: | ELECTRON MICROSCOPY (2.91 Å) | | Cite: | Mechanism of 30S subunit recognition and modification by the conserved bacterial ribosomal RNA methyltransferase RsmI.
Proc.Natl.Acad.Sci.USA, 123, 2026
|
|
10HY
 
 | | Structure of CHK1 10-pt. mutant complex with macrocyclic LRRK2 inhibitor compound 1 ((11R)-8-chloro-3,11-dimethyl-2-(oxan-4-yl)-2,4,10,11,12,13-hexahydro-9,5-(azeno)pyrazolo[3,4-b][1,4,6,10]oxatriazacyclotridecine) | | Descriptor: | (11R)-8-chloro-3,11-dimethyl-2-(oxan-4-yl)-2,4,10,11,12,13-hexahydro-9,5-(azeno)pyrazolo[3,4-b][1,4,6,10]oxatriazacyclotridecine, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L, Yu, E.C, Zhou, H. | | Deposit date: | 2026-01-21 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-22 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Discovery of Potent, Selective, CNS-Penetrant Macrocyclic LRRK2 Inhibitors for the Treatment of Parkinson's Disease.
J.Med.Chem., 69, 2026
|
|
10HZ
 
 | | Structure of CHK1 10-pt. mutant complex with macrocyclic LRRK2 inhibitor compound 7 ((10aS,13aS)-3-cyclobutyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine) | | Descriptor: | (10aS,13aS)-3-cyclobutyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L, Yu, E.C, Zhou, H. | | Deposit date: | 2026-01-21 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-22 | | Method: | X-RAY DIFFRACTION (1.666 Å) | | Cite: | Discovery of Potent, Selective, CNS-Penetrant Macrocyclic LRRK2 Inhibitors for the Treatment of Parkinson's Disease.
J.Med.Chem., 69, 2026
|
|
10IA
 
 | | Structure of CHK1 10-pt. mutant complex with macrocyclic LRRK2 inhibitor compound 12 ((10aS,13aS)-3-cyclopropyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine) | | Descriptor: | (10aS,13aS)-3-cyclopropyl-1-methyl-8-(trifluoromethyl)-3,4,10a,11,13a,14-hexahydro-10H,13H-9,5-(azeno)furo[3,4-k]pyrazolo[4,3-b][1,4,6,10]oxatriazacyclotridecine, Serine/threonine-protein kinase Chk1 | | Authors: | Palte, R.L, Yu, E.C, Zhou, H. | | Deposit date: | 2026-01-21 | | Release date: | 2026-04-08 | | Last modified: | 2026-04-22 | | Method: | X-RAY DIFFRACTION (1.739 Å) | | Cite: | Discovery of Potent, Selective, CNS-Penetrant Macrocyclic LRRK2 Inhibitors for the Treatment of Parkinson's Disease.
J.Med.Chem., 69, 2026
|
|
10IC
 
 | | Rhesus rotavirus (consensus structure at 4.7 Angstrom resolution from cryo-ET) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, CHLORIDE ION, ... | | Authors: | de Sautu, M, Leistner, C, Kirchhausen, T, Jenni, S, Harrison, S.C. | | Deposit date: | 2026-01-21 | | Release date: | 2026-03-04 | | Method: | ELECTRON MICROSCOPY (4.7 Å) | | Cite: | Mechanism of membrane perforation in rotavirus cell entry.
Biorxiv, 2026
|
|
10JT
 
 | | CRYSTAL STRUCTURE OF KIRSTEN RAT SARCOMA G12C COMPLEXED WITH GMPPNP AND COVALENTLY BOUND TO 1-[(2R,3R)-3-{[(7P)-7-(8-ethynyl-7-fluoronaphthalen-1-yl)-8-fluoro-2-{ [(2R,4R,7aS)-2-fluorotetrahydro-1H-pyrrolizin-7a(5H)-yl]methoxy}pyrido[4,3-d] pyrimidin-4-yl](methyl)amino}-2-methylpyrrolidin-1-yl]-3-(pyrazin-2-yl)propan-1-one | | Descriptor: | 1-[(2R,3R)-3-{[(7P)-7-(8-ethynyl-7-fluoronaphthalen-1-yl)-8-fluoro-2-{[(2R,4R,7aS)-2-fluorotetrahydro-1H-pyrrolizin-7a(5H)-yl]methoxy}pyrido[4,3-d]pyrimidin-4-yl](methyl)amino}-2-methylpyrrolidin-1-yl]-3-(pyrazin-2-yl)propan-1-one, CALCIUM ION, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Sheriff, S. | | Deposit date: | 2026-01-22 | | Release date: | 2026-03-04 | | Last modified: | 2026-03-18 | | Method: | X-RAY DIFFRACTION (1.489 Å) | | Cite: | Optimization of Covalent Warhead Trajectory for KRAS G12C Active-State Inhibition.
J.Med.Chem., 69, 2026
|
|
10KZ
 
 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors | | Descriptor: | (3'R)-1'-(1-benzyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, Plasma kallikrein | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-26 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10LR
 
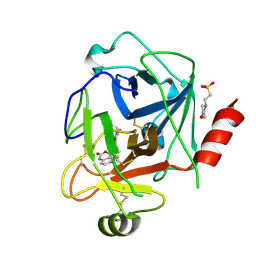 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable PlasmaKallikrein Inhibitors complex with Compound 4 ((3'R)-1'-(5-amino-1-benzyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-(5-amino-1-benzyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-27 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.583 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10MV
 
 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors complex with Compound 15 ((3'R)-1'-(5-amino-1-phenyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-(5-amino-1-phenyl-1H-pyrazole-4-carbonyl)-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Ogawa, A, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-28 | | Release date: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10MW
 
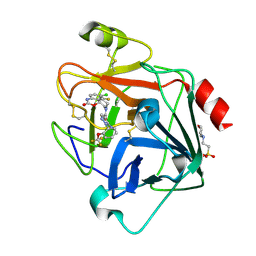 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors compound 25 ((3'R)-1'-{(1P)-5-amino-1-[2-(trifluoromethoxy)phenyl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-{(1P)-5-amino-1-[2-(trifluoromethoxy)phenyl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, Je, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yang, S, Cheng, A, Ellsworth, K, Poiou, T, Fier, P, Hicks, J, Sinz, C, Ogawa, A. | | Deposit date: | 2026-01-28 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10NU
 
 | | Structure of kRas G12C bound to Inhibitor 13ab | | Descriptor: | 1-((2R,5S)-4-((S)-6-chloro-7-(1,6-dimethyl-1H-indazol-7-yl)-8-fluoro-2-(((S)-1-methylpyrrolidin-2-yl)methoxy)quinazolin-4-yl)-2,5-dimethylpiperazin-1-yl)prop-2-en-1-one, CALCIUM ION, GLYCEROL, ... | | Authors: | Shaffer, P.L, Milligan, C, Peters, U. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Optimization of Covalent 6-Cyanoquinazoline KRAS G12C Inhibitors for the Treatment of Solid Tumors.
J.Med.Chem., 69, 2026
|
|
10NV
 
 | | Structure of kRas G12C Bound to Inhibitor 13ba | | Descriptor: | 4-((2S,5R)-4-Acryloyl-2,5-dimethylpiperazin-1-yl)-7-(1,6-dimethyl-1H-indazol-7-yl)-8-fluoro-2-(((S)-1-methylpyrrolidin-2-yl)methoxy)quinazoline-6-carbonitrile, CALCIUM ION, GTPase KRas, ... | | Authors: | Shaffer, P.L, Milligan, C. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Optimization of Covalent 6-Cyanoquinazoline KRAS G12C Inhibitors for the Treatment of Solid Tumors.
J.Med.Chem., 69, 2026
|
|
10OO
 
 | | FGFR2 mutant D650V with compound 4 (AZD3463) | | Descriptor: | (4P)-N-[4-(4-aminopiperidin-1-yl)-2-methoxyphenyl]-5-chloro-4-(1H-indol-3-yl)pyrimidin-2-amine, Fibroblast growth factor receptor 2, GLYCEROL, ... | | Authors: | Hoffman, I.D, Nelson, K.J. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure-Based Design of a Novel Covalent 4-(1-Methylindol-3-yl)pyrimidin-2-amine Series Targeting FGFR2 Resistance Mutations.
J.Med.Chem., 69, 2026
|
|
10OQ
 
 | | FGFR2 mutant D650V with compound 6 | | Descriptor: | Fibroblast growth factor receptor 2, GLYCEROL, N-[(3M)-3-{5-chloro-2-[4-(morpholin-4-yl)anilino]pyrimidin-4-yl}-1-methyl-1H-indol-6-yl]propanamide, ... | | Authors: | Hoffman, I.D, Nelson, K.J, Bensen, D.C, Rideout, M, Hudkins, R.L, Frye, C, Bailey, J.B. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structure-Based Design of a Novel Covalent 4-(1-Methylindol-3-yl)pyrimidin-2-amine Series Targeting FGFR2 Resistance Mutations.
J.Med.Chem., 69, 2026
|
|
10OU
 
 | | FGFR2 mutant D650V with compound 12 | | Descriptor: | 1,2-ETHANEDIOL, Fibroblast growth factor receptor 2, GLYCEROL, ... | | Authors: | Hoffman, I.D, Nelson, K.J, Bensen, D.C, Rideout, M, Hudkins, R.L, Frye, C, Bailey, J.B. | | Deposit date: | 2026-01-29 | | Release date: | 2026-04-15 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structure-Based Design of a Novel Covalent 4-(1-Methylindol-3-yl)pyrimidin-2-amine Series Targeting FGFR2 Resistance Mutations.
J.Med.Chem., 69, 2026
|
|
10PX
 
 | | Crystal structure of the wild-type Thermus thermophilus 70S ribosome in complex with benzoxaborole derivative of azithromycin (AZI-BB2), mRNA, aminoacylated A-site Phe-tRNAphe, aminoacylated P-site fMet-tRNAmet, and deacylated E-site tRNAphe at 2.45A resolution | | Descriptor: | 16S Ribosomal RNA, 23S Ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Chen, C.-W, Volynkina, I.A, Bortyazh, M.O, Tereshchenkov, A.G, Karakchieva, A.O, Lukianov, D.A, Komarova, E.S, Tupikin, A.E, Skvortsov, D.A, Tevyashova, A.N, Tikhomirov, A.S, Tashlitsky, V.N, Kabilov, M.R, Shchekotikhin, A.E, Dontsova, O.A, Sergiev, P.V, Polikanov, Y.S. | | Deposit date: | 2026-02-01 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-08 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Benzoxaborole-modified azithromycins inhibit translation without inducing ermC expression.
Antimicrob.Agents Chemother., 2026
|
|
10QS
 
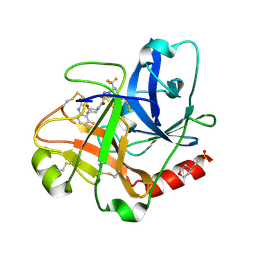 | | N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors Compound 13 ((3'R)-1'-{5-amino-1-[(2S)-1,1,1-trifluorobutan-2-yl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one) | | Descriptor: | (3'R)-1'-{5-amino-1-[(2S)-1,1,1-trifluorobutan-2-yl]-1H-pyrazole-4-carbonyl}-6-chloro-5-fluorospiro[[3,1]benzoxazine-4,3'-piperidin]-2(1H)-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Plasma kallikrein | | Authors: | Merchant, R.R, Chernyak, N, Lopez, J.A, Sharp, P.P, Mandal, M, He, J, Hruza, A, Rearden, P, Tatosian, D.A, Lin, K, Esmay, J, Yany, S, Cheng, A, Ellsworth, K, Piou, T, Fier, P, Hicks, J, Sinz, C, Ogowa, A. | | Deposit date: | 2026-02-02 | | Release date: | 2026-03-11 | | Last modified: | 2026-04-01 | | Method: | X-RAY DIFFRACTION (1.733 Å) | | Cite: | N ‐Alkyl and N ‐Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy to Identify Orally Bioavailable Plasma Kallikrein Inhibitors.
Acs Med.Chem.Lett., 17, 2026
|
|
10RA
 
 | | Human SARM1 TIR domain bound to compound 6 | | Descriptor: | 1-[(2R,3R,4S,5R)-5-({[(S)-{[(R)-{[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}methyl)-3,4-dihydroxyoxolan-2-yl]-4-[(4S,6P)-6-(5-fluoro-1H-indol-2-yl)imidazo[1,2-a]pyrazin-2-yl]pyridin-1-ium (non-preferred name), Sterile alpha and TIR motif-containing protein 1 | | Authors: | Olland, A.M, Johnson, Z.L, Nomme, J, Ciesielski, F, Ronin, C. | | Deposit date: | 2026-02-02 | | Release date: | 2026-04-22 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structure-Based Discovery of Imidazo[4,5- c ]pyridine SARM1 Modulators Showing Paradoxical Activation.
J.Med.Chem., 69, 2026
|
|
10RB
 
 | | Human SARM1 TIR domain bound to compound 14 | | Descriptor: | 1-[(2R,3R,4S,5R)-5-({[(S)-{[(S)-{[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}methyl)-3,4-dihydroxyoxolan-2-yl]-4-[(6P)-7-{[2-(dimethylamino)ethyl]amino}-6-(5-fluoro-1H-indol-2-yl)-1-methyl-1H-imidazo[4,5-c]pyridin-2-yl]pyridin-1-ium (non-preferred name), NAD(+) hydrolase SARM1 | | Authors: | Dementiev, A, Olland, A.M, Johnson, Z.L. | | Deposit date: | 2026-02-02 | | Release date: | 2026-04-22 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure-Based Discovery of Imidazo[4,5- c ]pyridine SARM1 Modulators Showing Paradoxical Activation.
J.Med.Chem., 69, 2026
|
|
10RC
 
 | | Human SARM1 TIR domain bound to compound 22 | | Descriptor: | 1-[(2R,3R,4S,5R)-5-({[(S)-{[(S)-{[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl]oxy}(hydroxy)phosphoryl]oxy}methyl)-3,4-dihydroxyoxolan-2-yl]-4-{(6P)-7-{[2-(dimethylamino)ethyl]amino}-6-[6-fluoro-4-(pyrimidin-2-yl)-1H-indol-2-yl]-1-methyl-1H-imidazo[4,5-c]pyridin-2-yl}pyridin-1-ium (non-preferred name), GLYCEROL, NAD(+) hydrolase SARM1 | | Authors: | Dementiev, A, Johnson, Z.L, Olland, A.M. | | Deposit date: | 2026-02-02 | | Release date: | 2026-04-22 | | Last modified: | 2026-04-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-Based Discovery of Imidazo[4,5- c ]pyridine SARM1 Modulators Showing Paradoxical Activation.
J.Med.Chem., 69, 2026
|
|
11AD
 
 | |
11BQ
 
 | |
11DG
 
 | |







