Entry Database : PDB / ID : 5i4zTitle Structure of apo OmoMYC Myc proto-oncogene protein Keywords / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / / / Resolution : 1.95 Å Authors Koelmel, W. / Jung, L.A. / Kuper, J. / Eilers, M. / Kisker, C. Funding support Organization Grant number Country German Research Foundation 2341
Journal : Oncogene / Year : 2017Title : OmoMYC blunts promoter invasion by oncogenic MYC to inhibit gene expression characteristic of MYC-dependent tumors.Authors: Jung, L.A. / Gebhardt, A. / Koelmel, W. / Ade, C.P. / Walz, S. / Kuper, J. / von Eyss, B. / Letschert, S. / Redel, C. / d'Artista, L. / Biankin, A. / Zender, L. / Sauer, M. / Wolf, E. / ... Authors : Jung, L.A. / Gebhardt, A. / Koelmel, W. / Ade, C.P. / Walz, S. / Kuper, J. / von Eyss, B. / Letschert, S. / Redel, C. / d'Artista, L. / Biankin, A. / Zender, L. / Sauer, M. / Wolf, E. / Evan, G. / Kisker, C. / Eilers, M. History Deposition Feb 13, 2016 Deposition site / Processing site Revision 1.0 Oct 26, 2016 Provider / Type Revision 1.1 Apr 19, 2017 Group Revision 1.2 Jan 10, 2024 Group Author supporting evidence / Data collection ... Author supporting evidence / Data collection / Database references / Derived calculations / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_audit_support / pdbx_initial_refinement_model / pdbx_struct_conn_angle / struct_conn Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_audit_support.funding_organization / _pdbx_struct_conn_angle.ptnr1_auth_asym_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr1_symmetry / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.ptnr3_symmetry / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_symmetry
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords TRANSCRIPTION /
TRANSCRIPTION /  leucine zipper /
leucine zipper /  transcription factor /
transcription factor /  tumor suppressor /
tumor suppressor /  E-box
E-box Function and homology information
Function and homology information SCF ubiquitin ligase complex binding / positive regulation of metanephric cap mesenchymal cell proliferation / negative regulation of transcription initiation by RNA polymerase II / Myc-Max complex / RNA polymerase II transcription repressor complex / regulation of cell cycle process / regulation of somatic stem cell population maintenance / Binding of TCF/LEF:CTNNB1 to target gene promoters / RUNX3 regulates WNT signaling / TFAP2 (AP-2) family regulates transcription of cell cycle factors ...
SCF ubiquitin ligase complex binding / positive regulation of metanephric cap mesenchymal cell proliferation / negative regulation of transcription initiation by RNA polymerase II / Myc-Max complex / RNA polymerase II transcription repressor complex / regulation of cell cycle process / regulation of somatic stem cell population maintenance / Binding of TCF/LEF:CTNNB1 to target gene promoters / RUNX3 regulates WNT signaling / TFAP2 (AP-2) family regulates transcription of cell cycle factors ... SCF ubiquitin ligase complex binding / positive regulation of metanephric cap mesenchymal cell proliferation / negative regulation of transcription initiation by RNA polymerase II / Myc-Max complex / RNA polymerase II transcription repressor complex / regulation of cell cycle process / regulation of somatic stem cell population maintenance / Binding of TCF/LEF:CTNNB1 to target gene promoters / RUNX3 regulates WNT signaling / TFAP2 (AP-2) family regulates transcription of cell cycle factors / negative regulation of cell division / negative regulation of monocyte differentiation / DNA methylation-dependent heterochromatin formation / Transcription of E2F targets under negative control by DREAM complex / protein-DNA complex disassembly / transcription regulator activator activity / response to growth factor / negative regulation of stress-activated MAPK cascade /
SCF ubiquitin ligase complex binding / positive regulation of metanephric cap mesenchymal cell proliferation / negative regulation of transcription initiation by RNA polymerase II / Myc-Max complex / RNA polymerase II transcription repressor complex / regulation of cell cycle process / regulation of somatic stem cell population maintenance / Binding of TCF/LEF:CTNNB1 to target gene promoters / RUNX3 regulates WNT signaling / TFAP2 (AP-2) family regulates transcription of cell cycle factors / negative regulation of cell division / negative regulation of monocyte differentiation / DNA methylation-dependent heterochromatin formation / Transcription of E2F targets under negative control by DREAM complex / protein-DNA complex disassembly / transcription regulator activator activity / response to growth factor / negative regulation of stress-activated MAPK cascade /  regulation of telomere maintenance / fibroblast apoptotic process / Regulation of NFE2L2 gene expression / positive regulation of mesenchymal cell proliferation / branching involved in ureteric bud morphogenesis / Signaling by ALK /
regulation of telomere maintenance / fibroblast apoptotic process / Regulation of NFE2L2 gene expression / positive regulation of mesenchymal cell proliferation / branching involved in ureteric bud morphogenesis / Signaling by ALK /  E-box binding / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / positive regulation of cysteine-type endopeptidase activity involved in apoptotic process / chromosome organization / Cyclin E associated events during G1/S transition / negative regulation of fibroblast proliferation / core promoter sequence-specific DNA binding / Cyclin A:Cdk2-associated events at S phase entry / positive regulation of telomerase activity / ERK1 and ERK2 cascade / positive regulation of epithelial cell proliferation /
E-box binding / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / positive regulation of cysteine-type endopeptidase activity involved in apoptotic process / chromosome organization / Cyclin E associated events during G1/S transition / negative regulation of fibroblast proliferation / core promoter sequence-specific DNA binding / Cyclin A:Cdk2-associated events at S phase entry / positive regulation of telomerase activity / ERK1 and ERK2 cascade / positive regulation of epithelial cell proliferation /  transcription coregulator binding / response to gamma radiation / SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription / G1/S transition of mitotic cell cycle / MAPK6/MAPK4 signaling / NOTCH1 Intracellular Domain Regulates Transcription / Constitutive Signaling by NOTCH1 PEST Domain Mutants / Constitutive Signaling by NOTCH1 HD+PEST Domain Mutants / DNA-binding transcription repressor activity, RNA polymerase II-specific / positive regulation of miRNA transcription / Transcriptional regulation of granulopoiesis / cellular response to UV / positive regulation of fibroblast proliferation /
transcription coregulator binding / response to gamma radiation / SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription / G1/S transition of mitotic cell cycle / MAPK6/MAPK4 signaling / NOTCH1 Intracellular Domain Regulates Transcription / Constitutive Signaling by NOTCH1 PEST Domain Mutants / Constitutive Signaling by NOTCH1 HD+PEST Domain Mutants / DNA-binding transcription repressor activity, RNA polymerase II-specific / positive regulation of miRNA transcription / Transcriptional regulation of granulopoiesis / cellular response to UV / positive regulation of fibroblast proliferation /  MAPK cascade / cellular response to xenobiotic stimulus / cellular response to hypoxia /
MAPK cascade / cellular response to xenobiotic stimulus / cellular response to hypoxia /  regulation of gene expression / DNA-binding transcription activator activity, RNA polymerase II-specific / Interleukin-4 and Interleukin-13 signaling / DNA-binding transcription factor binding / Estrogen-dependent gene expression / intracellular iron ion homeostasis /
regulation of gene expression / DNA-binding transcription activator activity, RNA polymerase II-specific / Interleukin-4 and Interleukin-13 signaling / DNA-binding transcription factor binding / Estrogen-dependent gene expression / intracellular iron ion homeostasis /  protein dimerization activity / DNA-binding transcription factor activity, RNA polymerase II-specific / Ub-specific processing proteases /
protein dimerization activity / DNA-binding transcription factor activity, RNA polymerase II-specific / Ub-specific processing proteases /  chromatin remodeling / response to xenobiotic stimulus / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA damage response / positive regulation of cell population proliferation /
chromatin remodeling / response to xenobiotic stimulus / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA damage response / positive regulation of cell population proliferation /  chromatin / protein-containing complex binding / positive regulation of gene expression /
chromatin / protein-containing complex binding / positive regulation of gene expression /  nucleolus / regulation of transcription by RNA polymerase II / negative regulation of apoptotic process / positive regulation of DNA-templated transcription / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / protein-containing complex /
nucleolus / regulation of transcription by RNA polymerase II / negative regulation of apoptotic process / positive regulation of DNA-templated transcription / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / protein-containing complex /  DNA binding /
DNA binding /  nucleoplasm / identical protein binding /
nucleoplasm / identical protein binding /  nucleus
nucleus
 Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.95 Å
MOLECULAR REPLACEMENT / Resolution: 1.95 Å  Authors
Authors Germany, 1items
Germany, 1items  Citation
Citation Journal: Oncogene / Year: 2017
Journal: Oncogene / Year: 2017 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5i4z.cif.gz
5i4z.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5i4z.ent.gz
pdb5i4z.ent.gz PDB format
PDB format 5i4z.json.gz
5i4z.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/i4/5i4z
https://data.pdbj.org/pub/pdb/validation_reports/i4/5i4z ftp://data.pdbj.org/pub/pdb/validation_reports/i4/5i4z
ftp://data.pdbj.org/pub/pdb/validation_reports/i4/5i4z

 Links
Links Assembly
Assembly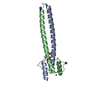
 Components
Components
 Homo sapiens (human) / Gene: MYC, BHLHE39 / Production host:
Homo sapiens (human) / Gene: MYC, BHLHE39 / Production host: 
 Escherichia coli (E. coli) / References: UniProt: P01106
Escherichia coli (E. coli) / References: UniProt: P01106 Glycerol
Glycerol Chloride
Chloride Water
Water X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation
 SYNCHROTRON / Site:
SYNCHROTRON / Site:  ESRF
ESRF  / Beamline: ID23-1 / Wavelength: 1.064 Å
/ Beamline: ID23-1 / Wavelength: 1.064 Å : 1.064 Å / Relative weight: 1
: 1.064 Å / Relative weight: 1  Processing
Processing :
:  MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller













 PDBj
PDBj


















