+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3dxb | ||||||
|---|---|---|---|---|---|---|---|
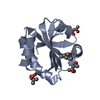
| Title | Structure of the UHM domain of Puf60 fused to thioredoxin | ||||||
 Components Components | thioredoxin N-terminally fused to Puf60(UHM) | ||||||
 Keywords Keywords | SPLICING / TRANSCRIPTION / FBP interacting repressor / UHM / RRM / Electron transport / Redox-active center / Transport | ||||||
| Function / homology |  Function and homology information Function and homology informationmRNA splice site recognition / alternative mRNA splicing, via spliceosome / regulation of alternative mRNA splicing, via spliceosome / protein-disulfide reductase activity / mRNA Splicing - Major Pathway / cell redox homeostasis / cell junction / cadherin binding / ribonucleoprotein complex / apoptotic process ...mRNA splice site recognition / alternative mRNA splicing, via spliceosome / regulation of alternative mRNA splicing, via spliceosome / protein-disulfide reductase activity / mRNA Splicing - Major Pathway / cell redox homeostasis / cell junction / cadherin binding / ribonucleoprotein complex / apoptotic process / DNA binding / RNA binding / nucleoplasm / identical protein binding / cytosol Similarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Corsini, L. / Hothorn, M. / Scheffzek, K. / Stier, G. / Sattler, M. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2009 Journal: J.Biol.Chem. / Year: 2009Title: Dimerization and Protein Binding Specificity of the U2AF Homology Motif of the Splicing Factor Puf60. Authors: Corsini, L. / Hothorn, M. / Stier, G. / Rybin, V. / Scheffzek, K. / Gibson, T.J. / Sattler, M. #1: Journal: Protein Sci. / Year: 2008 Title: Thioredoxin as a fusion tag for carrier-driven crystallization. Authors: Corsini, L. / Hothorn, M. / Scheffzek, K. / Sattler, M. / Stier, G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3dxb.cif.gz 3dxb.cif.gz | 349.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3dxb.ent.gz pdb3dxb.ent.gz | 283.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3dxb.json.gz 3dxb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dx/3dxb https://data.pdbj.org/pub/pdb/validation_reports/dx/3dxb ftp://data.pdbj.org/pub/pdb/validation_reports/dx/3dxb ftp://data.pdbj.org/pub/pdb/validation_reports/dx/3dxb | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| 5 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 24671.930 Da / Num. of mol.: 8 Fragment: Chimera of Thioredoxin 1-109 and Puf60 C-terminal 460-559 Source method: isolated from a genetically manipulated source Details: The UHM domain of Puf60 was crystallized as a fusion with E.coli thioredoxin (U niProtKB/Swiss-Prot P0AA27) Source: (gene. exp.)   Homo sapiens (human) Homo sapiens (human)Gene: trxA, Z5291, ECs4714, PUF60,FIR,ROBPI,SIAHBP1 / Plasmid: pET9d / Production host:  #2: Chemical | ChemComp-CL / #3: Chemical | ChemComp-EDO / | #4: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.55 Å3/Da / Density % sol: 51.72 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 7 Details: 1.4M ammonium sulfate, 0.05M K-formate, pH 7.0, VAPOR DIFFUSION, SITTING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X06SA / Wavelength: 1.006 Å / Beamline: X06SA / Wavelength: 1.006 Å |
| Detector | Type: MARMOSAIC 225 mm CCD / Detector: CCD / Date: May 6, 2007 Details: LN2 cooled fixed-exit Si(111) monochromator Dynamically bendable mirror Beamline suitable for large unit cell structures |
| Radiation | Monochromator: LN2 cooled fixed-exit Si(111) monochromator / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.006 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→49.832 Å / Num. all: 103128 / Num. obs: 102916 / % possible obs: 99.8 % / Observed criterion σ(F): 1 / Observed criterion σ(I): 1 / Redundancy: 6.1 % / Biso Wilson estimate: 36.481 Å2 / Rmerge(I) obs: 0.11 / Rsym value: 0.106 / Net I/σ(I): 14.53 |
| Reflection shell | Resolution: 2.2→2.33 Å / Redundancy: 5.8 % / Rmerge(I) obs: 0.393 / Mean I/σ(I) obs: 3.24 / Num. unique all: 16266 / Rsym value: 0.604 / % possible all: 98.9 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 8 thioredoxin domains: PDB entry 2TRX 8 Puf60-UHM domains: homology model based on PDB 2pe8 Resolution: 2.2→49.75 Å / Cor.coef. Fo:Fc: 0.938 / Cor.coef. Fo:Fc free: 0.896 / SU B: 12.772 / SU ML: 0.185 / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 2 / ESU R: 0.282 / ESU R Free: 0.233 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 21.623 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→49.75 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.2→2.259 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller














 PDBj
PDBj









