+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ify | ||||||
|---|---|---|---|---|---|---|---|
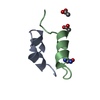
| Title | Solution Structure of the Internal UBA Domain of HHR23A | ||||||
 Components Components | UV EXCISION REPAIR PROTEIN RAD23 HOMOLOG A | ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / Ubiquitin associated domain / DNA BINDING PROTEIN / Ubiquitin associated domain /  UBA domain / Ubiquitin proteosome pathway UBA domain / Ubiquitin proteosome pathway | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of proteasomal ubiquitin-dependent protein catabolic process /  proteasome binding / ubiquitin-specific protease binding / polyubiquitin modification-dependent protein binding / positive regulation of viral genome replication / positive regulation of cell cycle / proteasome binding / ubiquitin-specific protease binding / polyubiquitin modification-dependent protein binding / positive regulation of viral genome replication / positive regulation of cell cycle /  proteasome complex / Josephin domain DUBs / proteasome complex / Josephin domain DUBs /  ubiquitin binding / nucleotide-excision repair ...regulation of proteasomal ubiquitin-dependent protein catabolic process / ubiquitin binding / nucleotide-excision repair ...regulation of proteasomal ubiquitin-dependent protein catabolic process /  proteasome binding / ubiquitin-specific protease binding / polyubiquitin modification-dependent protein binding / positive regulation of viral genome replication / positive regulation of cell cycle / proteasome binding / ubiquitin-specific protease binding / polyubiquitin modification-dependent protein binding / positive regulation of viral genome replication / positive regulation of cell cycle /  proteasome complex / Josephin domain DUBs / proteasome complex / Josephin domain DUBs /  ubiquitin binding / nucleotide-excision repair / DNA Damage Recognition in GG-NER / protein destabilization / Formation of Incision Complex in GG-NER / ubiquitin binding / nucleotide-excision repair / DNA Damage Recognition in GG-NER / protein destabilization / Formation of Incision Complex in GG-NER /  kinase binding / positive regulation of proteasomal ubiquitin-dependent protein catabolic process / kinase binding / positive regulation of proteasomal ubiquitin-dependent protein catabolic process /  single-stranded DNA binding / proteasome-mediated ubiquitin-dependent protein catabolic process / damaged DNA binding / intracellular membrane-bounded organelle / single-stranded DNA binding / proteasome-mediated ubiquitin-dependent protein catabolic process / damaged DNA binding / intracellular membrane-bounded organelle /  Golgi apparatus / protein-containing complex / Golgi apparatus / protein-containing complex /  nucleoplasm / nucleoplasm /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Mueller, T.D. / Feigon, J. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2002 Journal: J.Mol.Biol. / Year: 2002Title: Solution structures of UBA domains reveal a conserved hydrophobic surface for protein-protein interactions. Authors: Mueller, T.D. / Feigon, J. #1:  Journal: Biochemistry / Year: 2000 Journal: Biochemistry / Year: 2000Title: Biochemical and Structural Analysis of the Interaction between the Uba(2) Domain of the DNA Repair Protein Hhr23A and HIV-1 Vpr Authors: Withers-Ward, E.S. / Mueller, T.D. / Chen, I.S. / Feigon, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ify.cif.gz 1ify.cif.gz | 152.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ify.ent.gz pdb1ify.ent.gz | 129.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ify.json.gz 1ify.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/if/1ify https://data.pdbj.org/pub/pdb/validation_reports/if/1ify ftp://data.pdbj.org/pub/pdb/validation_reports/if/1ify ftp://data.pdbj.org/pub/pdb/validation_reports/if/1ify | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 5496.235 Da / Num. of mol.: 1 / Fragment: INTERNAL UBA DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Plasmid: pGEX2T / Production host: Homo sapiens (human) / Plasmid: pGEX2T / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3) Star / References: UniProt: P54725 Escherichia coli (E. coli) / Strain (production host): BL21(DE3) Star / References: UniProt: P54725 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: The structure was determined using triple-resonance NMR spectroscopy |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 50mM sodium phosphate, 100mM sodium chloride pH: 6.5 / Pressure: ambient / Temperature: 300 K | ||||||||||||
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Bruker DRX / Manufacturer: Bruker / Model : DRX / Field strength: 500 MHz : DRX / Field strength: 500 MHz |
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 Details: The structures are based on 1692 NOE-derived distance restraints, of which 1055 distance restraints are non-redundant. 407 restraints are intraresidue, 212 are sequential, 226 are ...Details: The structures are based on 1692 NOE-derived distance restraints, of which 1055 distance restraints are non-redundant. 407 restraints are intraresidue, 212 are sequential, 226 are mediumrange (i-j < 5) and 210 are long-range (i-j > 4) | ||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 100 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller














 PDBj
PDBj


