[English] 日本語
 Yorodumi
Yorodumi- PDB-6y4s: Human kallikrein-related peptidase 7 (KLK7) in the unliganded state -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6y4s | ||||||
|---|---|---|---|---|---|---|---|
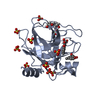
| Title | Human kallikrein-related peptidase 7 (KLK7) in the unliganded state | ||||||
 Components Components | Kallikrein-7 | ||||||
 Keywords Keywords | HYDROLASE / kallikrein 7 / hK7 / serine protease | ||||||
| Function / homology |  Function and homology information Function and homology informationstratum corneum chymotryptic enzyme / positive regulation of antibacterial peptide production / epidermal lamellar body / cornified envelope / Differentiation of Keratinocytes in Interfollicular Epidermis in Mammalian Skin / extracellular matrix disassembly / epidermis development / Degradation of the extracellular matrix / serine-type peptidase activity / secretory granule ...stratum corneum chymotryptic enzyme / positive regulation of antibacterial peptide production / epidermal lamellar body / cornified envelope / Differentiation of Keratinocytes in Interfollicular Epidermis in Mammalian Skin / extracellular matrix disassembly / epidermis development / Degradation of the extracellular matrix / serine-type peptidase activity / secretory granule / protein maturation / metalloendopeptidase activity / peptidase activity / serine-type endopeptidase activity / proteolysis / extracellular space / extracellular region Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  FOURIER SYNTHESIS / Resolution: 2.23 Å FOURIER SYNTHESIS / Resolution: 2.23 Å | ||||||
| Model details | The compound is a competitive non-nucleotide inhibitor binding to the active site | ||||||
 Authors Authors | Hanke, S. / Strater, N. | ||||||
 Citation Citation |  Journal: J.Med.Chem. / Year: 2020 Journal: J.Med.Chem. / Year: 2020Title: Structural Studies on the Inhibitory Binding Mode of Aromatic Coumarinic Esters to Human Kallikrein-Related Peptidase 7. Authors: Hanke, S. / Tindall, C.A. / Pippel, J. / Ulbricht, D. / Pirotte, B. / Reboud-Ravaux, M. / Heiker, J.T. / Strater, N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6y4s.cif.gz 6y4s.cif.gz | 272.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6y4s.ent.gz pdb6y4s.ent.gz | 221.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6y4s.json.gz 6y4s.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  6y4s_validation.pdf.gz 6y4s_validation.pdf.gz | 462 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  6y4s_full_validation.pdf.gz 6y4s_full_validation.pdf.gz | 466.6 KB | Display | |
| Data in XML |  6y4s_validation.xml.gz 6y4s_validation.xml.gz | 25.1 KB | Display | |
| Data in CIF |  6y4s_validation.cif.gz 6y4s_validation.cif.gz | 34.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/y4/6y4s https://data.pdbj.org/pub/pdb/validation_reports/y4/6y4s ftp://data.pdbj.org/pub/pdb/validation_reports/y4/6y4s ftp://data.pdbj.org/pub/pdb/validation_reports/y4/6y4s | HTTPS FTP |
-Related structure data
| Related structure data |  6shhC 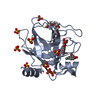 6shiC  6sjuC  3bsqS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 24481.160 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: KLK7, PRSS6, SCCE / Production host: Homo sapiens (human) / Gene: KLK7, PRSS6, SCCE / Production host:  References: UniProt: P49862, stratum corneum chymotryptic enzyme #2: Chemical | ChemComp-PGE / | #3: Chemical | ChemComp-SO4 / #4: Water | ChemComp-HOH / | Has ligand of interest | N | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.74 Å3/Da / Density % sol: 55.03 % |
|---|---|
| Crystal grow | Temperature: 292 K / Method: vapor diffusion, hanging drop / pH: 8.5 Details: 2.9 M ammonium sulphate, 0.1 M HEPES, pH 8.5 and 0.5-2 % PEG 3350 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  BESSY BESSY  / Beamline: 14.1 / Wavelength: 0.9184 Å / Beamline: 14.1 / Wavelength: 0.9184 Å | ||||||||||||||||||||||||||||||
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jan 23, 2018 | ||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.9184 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.23→48.45 Å / Num. obs: 39789 / % possible obs: 99.8 % / Redundancy: 15 % / Biso Wilson estimate: 40.47 Å2 / CC1/2: 0.994 / Rmerge(I) obs: 0.436 / Rpim(I) all: 0.116 / Rrim(I) all: 0.452 / Net I/σ(I): 6.3 / Num. measured all: 596786 | ||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  FOURIER SYNTHESIS FOURIER SYNTHESISStarting model: 3bsq Resolution: 2.23→48.45 Å / Cor.coef. Fo:Fc: 0.915 / Cor.coef. Fo:Fc free: 0.9 / SU R Cruickshank DPI: 0.258 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.259 / SU Rfree Blow DPI: 0.194 / SU Rfree Cruickshank DPI: 0.195
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 172.92 Å2 / Biso mean: 56.26 Å2 / Biso min: 18.06 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.41 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.23→48.45 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.23→2.24 Å / Rfactor Rfree error: 0 / Total num. of bins used: 50
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj






