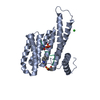
Entry Database : PDB / ID : 6qdtTitle Crystal structure of 14-3-3sigma in complex with a RapGef2 pT740 phosphopeptide 14-3-3 protein sigma Rap guanine nucleotide exchange factor 2 Keywords / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / / / Resolution : 1.702 Å Authors Andrei, S.A. / Kaplan, A. / Fournier, A.E. / Ottman, C. Funding support Organization Grant number Country Netherlands Organisation for Scientific Research ECHO -STIP 717.014.001
Journal : Cell Chem Biol / Year : 2020Title : Polypharmacological Perturbation of the 14-3-3 Adaptor Protein Interactome Stimulates Neurite Outgrowth.Authors : Kaplan, A. / Andrei, S.A. / van Regteren Altena, A. / Simas, T. / Banerjee, S.L. / Kato, N. / Bisson, N. / Higuchi, Y. / Ottmann, C. / Fournier, A.E. History Deposition Jan 2, 2019 Deposition site / Processing site Revision 1.0 Jan 29, 2020 Provider / Type Revision 1.1 Aug 11, 2021 Group / Derived calculationsCategory citation / citation_author ... citation / citation_author / database_2 / pdbx_struct_conn_angle / struct_conn / struct_conn_type Item _citation.country / _citation.journal_abbrev ... _citation.country / _citation.journal_abbrev / _citation.journal_id_CSD / _citation.journal_id_ISSN / _citation.journal_volume / _citation.page_first / _citation.page_last / _citation.pdbx_database_id_DOI / _citation.pdbx_database_id_PubMed / _citation.title / _citation.year / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_alt_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr1_symmetry / _pdbx_struct_conn_angle.ptnr2_auth_comp_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr2_label_atom_id / _pdbx_struct_conn_angle.ptnr2_label_comp_id / _pdbx_struct_conn_angle.ptnr2_symmetry / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_alt_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.ptnr3_symmetry / _pdbx_struct_conn_angle.value / _struct_conn.conn_type_id / _struct_conn.id / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.pdbx_ptnr1_label_alt_id / _struct_conn.pdbx_ptnr2_label_alt_id / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_conn.ptnr2_symmetry / _struct_conn_type.id Revision 1.2 Jan 24, 2024 Group / Refinement descriptionCategory / chem_comp_bond / pdbx_initial_refinement_modelRevision 1.3 Nov 6, 2024 Group / Category / pdbx_modification_feature
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT /
MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.702 Å
molecular replacement / Resolution: 1.702 Å  Authors
Authors Netherlands, 1items
Netherlands, 1items  Citation
Citation Journal: Cell Chem Biol / Year: 2020
Journal: Cell Chem Biol / Year: 2020 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6qdt.cif.gz
6qdt.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6qdt.ent.gz
pdb6qdt.ent.gz PDB format
PDB format 6qdt.json.gz
6qdt.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/qd/6qdt
https://data.pdbj.org/pub/pdb/validation_reports/qd/6qdt ftp://data.pdbj.org/pub/pdb/validation_reports/qd/6qdt
ftp://data.pdbj.org/pub/pdb/validation_reports/qd/6qdt



 Links
Links Assembly
Assembly

 Components
Components Homo sapiens (human) / Gene: SFN, HME1 / Production host:
Homo sapiens (human) / Gene: SFN, HME1 / Production host: 
 Homo sapiens (human) / References: UniProt: Q9Y4G8
Homo sapiens (human) / References: UniProt: Q9Y4G8






 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation molecular replacement
molecular replacement Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller
















 PDBj
PDBj


















