[English] 日本語
 Yorodumi
Yorodumi- PDB-6mae: CHAIN A. UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacety... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6mae | ||||||
|---|---|---|---|---|---|---|---|
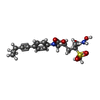
| Title | CHAIN A. UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase PA-LPXC Complexed with (R)-3-((S)-3-(4-(cyclopropylethynyl)phenyl)-2-oxooxazolidin-5-yl)-N-hydroxy-2-methyl-2-(methylsulfonyl)propenamide | ||||||
 Components Components | UDP-3-O-acyl-N-acetylglucosamine deacetylase | ||||||
 Keywords Keywords | HYDROLASE / LPXC | ||||||
| Function / homology |  Function and homology information Function and homology informationUDP-3-O-acyl-N-acetylglucosamine deacetylase / UDP-3-O-acyl-N-acetylglucosamine deacetylase activity / lipid A biosynthetic process / membrane / metal ion binding / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.8 Å molecular replacement / Resolution: 1.8 Å | ||||||
 Authors Authors | Shu, W. | ||||||
 Citation Citation |  Journal: J. Med. Chem. / Year: 2018 Journal: J. Med. Chem. / Year: 2018Title: Application of Virtual Screening to the Identification of New LpxC Inhibitor Chemotypes, Oxazolidinone and Isoxazoline. Authors: Lee, P.S. / Lapointe, G. / Madera, A.M. / Simmons, R.L. / Xu, W. / Yifru, A. / Tjandra, M. / Karur, S. / Rico, A. / Thompson, K. / Bojkovic, J. / Xie, L. / Uehara, K. / Liu, A. / Shu, W. / ...Authors: Lee, P.S. / Lapointe, G. / Madera, A.M. / Simmons, R.L. / Xu, W. / Yifru, A. / Tjandra, M. / Karur, S. / Rico, A. / Thompson, K. / Bojkovic, J. / Xie, L. / Uehara, K. / Liu, A. / Shu, W. / Bellamacina, C. / McKenney, D. / Morris, L. / Tonn, G.R. / Osborne, C. / Benton, B.M. / McDowell, L. / Fu, J. / Sweeney, Z.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6mae.cif.gz 6mae.cif.gz | 136.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6mae.ent.gz pdb6mae.ent.gz | 104.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6mae.json.gz 6mae.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ma/6mae https://data.pdbj.org/pub/pdb/validation_reports/ma/6mae ftp://data.pdbj.org/pub/pdb/validation_reports/ma/6mae ftp://data.pdbj.org/pub/pdb/validation_reports/ma/6mae | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 33162.680 Da / Num. of mol.: 1 / Source method: isolated from a natural source Source: (natural)  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 References: UniProt: P47205, UDP-3-O-acyl-N-acetylglucosamine deacetylase |
|---|---|
| #2: Chemical | ChemComp-JBA / ( |
| #3: Chemical | ChemComp-ZN / |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.3 Å3/Da / Density % sol: 46.47 % |
|---|---|
| Crystal grow | Temperature: 289 K / Method: vapor diffusion, hanging drop Details: 0.1 M MES PH 6.5, 20% PEG4000, 0.6M SODIUM CHLORIDE |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALS ALS  / Beamline: 5.0.2 / Wavelength: 1.00003 Å / Beamline: 5.0.2 / Wavelength: 1.00003 Å | ||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Aug 15, 2013 | ||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.00003 Å / Relative weight: 1 | ||||||||||||||||||||||||
| Reflection | Resolution: 1.8→67.35 Å / Num. obs: 27108 / % possible obs: 97.8 % / Redundancy: 3.8 % / Biso Wilson estimate: 19.25 Å2 / CC1/2: 0.997 / Rmerge(I) obs: 0.095 / Rpim(I) all: 0.057 / Rrim(I) all: 0.111 / Net I/σ(I): 11.9 | ||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement |
|---|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.8→62.86 Å / Cor.coef. Fo:Fc: 0.958 / Cor.coef. Fo:Fc free: 0.9416 / SU R Cruickshank DPI: 0.117 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.124 / SU Rfree Blow DPI: 0.117 / SU Rfree Cruickshank DPI: 0.114 MOLECULAR REPLACEMENT / Resolution: 1.8→62.86 Å / Cor.coef. Fo:Fc: 0.958 / Cor.coef. Fo:Fc free: 0.9416 / SU R Cruickshank DPI: 0.117 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.124 / SU Rfree Blow DPI: 0.117 / SU Rfree Cruickshank DPI: 0.114
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 96.47 Å2 / Biso mean: 21.67 Å2 / Biso min: 4.57 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.172 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.8→62.86 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.8→1.87 Å / Rfactor Rfree error: 0 / Total num. of bins used: 14
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 10.9274 Å / Origin y: 0.8353 Å / Origin z: 12.7267 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: { A|* } |
 Movie
Movie Controller
Controller






















 PDBj
PDBj