[English] 日本語
 Yorodumi
Yorodumi- PDB-6bbp: Model for compact volume of truncated monomeric Cytohesin-3 (Grp1... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6bbp | ||||||
|---|---|---|---|---|---|---|---|
| Title | Model for compact volume of truncated monomeric Cytohesin-3 (Grp1; amino acids 63-399) E161A 6GS Arf6 Q67L fusion protein | ||||||
 Components Components | Cytohesin-3,ADP-ribosylation factor 6 | ||||||
 Keywords Keywords | LIPID BINDING PROTEIN / Guanine nucleotide exchange factor / Arf GTPase / Fusion protein / Inositol 1 / 3 / 4 / 5-tetrakisphosphate | ||||||
| Function / homology |  Function and homology information Function and homology informationerythrocyte apoptotic process / maintenance of postsynaptic density structure / Intra-Golgi traffic / protein localization to cleavage furrow / positive regulation of mitotic cytokinetic process / establishment of epithelial cell polarity / Golgi vesicle transport / regulation of dendritic spine development / negative regulation of protein localization to cell surface / protein localization to endosome ...erythrocyte apoptotic process / maintenance of postsynaptic density structure / Intra-Golgi traffic / protein localization to cleavage furrow / positive regulation of mitotic cytokinetic process / establishment of epithelial cell polarity / Golgi vesicle transport / regulation of dendritic spine development / negative regulation of protein localization to cell surface / protein localization to endosome / regulation of Rac protein signal transduction / negative regulation of dendrite development / ruffle assembly / regulation of filopodium assembly / positive regulation of keratinocyte migration / positive regulation of focal adhesion disassembly / endocytic recycling / MET receptor recycling / thioesterase binding / regulation of ARF protein signal transduction / Flemming body / filopodium membrane / TBC/RABGAPs / protein localization to cell surface / cortical actin cytoskeleton organization / hepatocyte apoptotic process / phosphatidylinositol-3,4,5-trisphosphate binding / positive regulation of actin filament polymerization / cleavage furrow / bicellular tight junction / endocytic vesicle / regulation of presynapse assembly / synaptic vesicle endocytosis / vesicle-mediated transport / ruffle / signaling adaptor activity / guanyl-nucleotide exchange factor activity / positive regulation of cell adhesion / positive regulation of protein secretion / small monomeric GTPase / protein localization to plasma membrane / positive regulation of protein localization to plasma membrane / adherens junction / liver development / negative regulation of receptor-mediated endocytosis / cellular response to nerve growth factor stimulus / intracellular protein transport / positive regulation of neuron projection development / recycling endosome membrane / GDP binding / nervous system development / Clathrin-mediated endocytosis / presynapse / G protein activity / midbody / early endosome membrane / cell cortex / cell differentiation / cell adhesion / endosome / postsynapse / cell division / focal adhesion / GTPase activity / GTP binding / glutamatergic synapse / Golgi apparatus / extracellular exosome / nucleoplasm / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / negative staining / Resolution: 35 Å | ||||||
 Authors Authors | Das, S. / Malaby, A.W. / Lambright, D.G. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Structure / Year: 2018 Journal: Structure / Year: 2018Title: Structural Dynamics Control Allosteric Activation of Cytohesin Family Arf GTPase Exchange Factors. Authors: Andrew W Malaby / Sanchaita Das / Srinivas Chakravarthy / Thomas C Irving / Osman Bilsel / David G Lambright /  Abstract: Membrane dynamic processes including vesicle biogenesis depend on Arf guanosine triphosphatase (GTPase) activation by guanine nucleotide exchange factors (GEFs) containing a catalytic Sec7 domain and ...Membrane dynamic processes including vesicle biogenesis depend on Arf guanosine triphosphatase (GTPase) activation by guanine nucleotide exchange factors (GEFs) containing a catalytic Sec7 domain and a membrane-targeting module such as a pleckstrin homology (PH) domain. The catalytic output of cytohesin family Arf GEFs is controlled by autoinhibitory interactions that impede accessibility of the exchange site in the Sec7 domain. These restraints can be relieved through activator Arf-GTP binding to an allosteric site comprising the PH domain and proximal autoinhibitory elements (Sec7-PH linker and C-terminal helix). Small-angle X-ray scattering and negative-stain electron microscopy were used to investigate the structural organization and conformational dynamics of cytohesin-3 (Grp1) in autoinhibited and active states. The results support a model in which hinge dynamics in the autoinhibited state expose the activator site for Arf-GTP binding, while subsequent C-terminal helix unlatching and repositioning unleash conformational entropy in the Sec7-PH linker to drive exposure of the exchange site. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6bbp.cif.gz 6bbp.cif.gz | 107.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6bbp.ent.gz pdb6bbp.ent.gz | 76.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6bbp.json.gz 6bbp.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bb/6bbp https://data.pdbj.org/pub/pdb/validation_reports/bb/6bbp ftp://data.pdbj.org/pub/pdb/validation_reports/bb/6bbp ftp://data.pdbj.org/pub/pdb/validation_reports/bb/6bbp | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7077MC  7078C  6bbqC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 60292.777 Da / Num. of mol.: 1 / Mutation: E161A,E161A Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) Homo sapiens (human)Gene: Cyth3, Grp1, Pscd3, ARF6 / Production host:  |
|---|---|
| #2: Chemical | ChemComp-GTP / |
| #3: Chemical | ChemComp-MG / |
| #4: Chemical | ChemComp-4IP / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Truncated monomeric Cytohesin-3 (Grp1; amino acids 63-399) E161A 6GS Arf6 Q67L fusion protein complex with GTP, Mg and Inositol 1,3,4,5 tetrakisphosphate Type: COMPLEX / Entity ID: #1 / Source: RECOMBINANT |
|---|---|
| Source (natural) | Organism:  |
| Source (recombinant) | Organism:  |
| Buffer solution | pH: 8 Details: 20 mM Tris, pH 8.0, 150 mM NaCl, 2 mM MgCl2, 0.1% 2-mercaptoethanol, and 0.001 mM IP4 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: YES / Vitrification applied: NO |
| EM staining | Type: NEGATIVE / Details: Stained with 0.75% (w/v) uranyl formate / Material: Uranyl Formate |
| Specimen support | Grid material: COPPER / Grid mesh size: 400 divisions/in. |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: FEI TECNAI 12 Details: Gatan Erlang Shen 785 camera used for collecting images |
|---|---|
| Electron gun | Electron source: LAB6 / Accelerating voltage: 120 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 60000 X / Nominal defocus max: 3200 nm / Nominal defocus min: 1200 nm / Cs: 2 mm |
| Image recording | Electron dose: 20 e/Å2 / Film or detector model: OTHER / Num. of grids imaged: 1 / Num. of real images: 369 Details: Gatan Erlang Shen 785 camera used for collecting images |
- Processing
Processing
| EM software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Image processing | Details: The images were X-ray corrected | ||||||||||||||||
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||
| Particle selection | Num. of particles selected: 10000 / Details: EMAN2 based manual particle picking | ||||||||||||||||
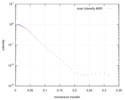
| 3D reconstruction | Resolution: 35 Å / Resolution method: FSC 0.5 CUT-OFF / Num. of particles: 6504 / Num. of class averages: 71 / Symmetry type: POINT | ||||||||||||||||
| Refinement | Highest resolution: 35 Å |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera















 PDBj
PDBj