Entry Database : PDB / ID : 5wleTitle Crystal structure of the PPS PHD finger in complex with H3K4me3 H3K4me3 Peptide Protein partner of snf, isoform A Keywords / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Drosophila melanogaster (fruit fly)Homo sapiens (human)Method / / Resolution : 1.952 Å Authors Klein, B.J. / Kutateladze, T.G. Funding support Organization Grant number Country National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) GM106416 National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) GM101664 National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) GM100907 National Institutes of Health/National Institute on Alcohol Abuse and Alcoholism (NIH/NIAAA) T32AA007464
Journal : Structure / Year : 2017Title : A Unique pH-Dependent Recognition of Methylated Histone H3K4 by PPS and DIDO.Authors : Tencer, A.H. / Gatchalian, J. / Klein, B.J. / Khan, A. / Zhang, Y. / Strahl, B.D. / van Wely, K.H.M. / Kutateladze, T.G. History Deposition Jul 26, 2017 Deposition site / Processing site Revision 1.0 Oct 4, 2017 Provider / Type Revision 1.1 Oct 18, 2017 Group / Category Item / _citation.page_first / _citation.page_lastRevision 1.2 Dec 11, 2019 Group / Category / Item Revision 1.3 Oct 4, 2023 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / struct_conn / struct_conn_type Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _struct_conn.conn_type_id / _struct_conn.id / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_conn_type.id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.952 Å
MOLECULAR REPLACEMENT / Resolution: 1.952 Å  Authors
Authors United States, 4items
United States, 4items  Citation
Citation Journal: Structure / Year: 2017
Journal: Structure / Year: 2017 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5wle.cif.gz
5wle.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5wle.ent.gz
pdb5wle.ent.gz PDB format
PDB format 5wle.json.gz
5wle.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/wl/5wle
https://data.pdbj.org/pub/pdb/validation_reports/wl/5wle ftp://data.pdbj.org/pub/pdb/validation_reports/wl/5wle
ftp://data.pdbj.org/pub/pdb/validation_reports/wl/5wle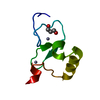

 Links
Links Assembly
Assembly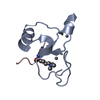
 Components
Components

 Homo sapiens (human) / References: UniProt: P68431*PLUS
Homo sapiens (human) / References: UniProt: P68431*PLUS X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU MICROMAX-007 / Wavelength: 1.54 Å
ROTATING ANODE / Type: RIGAKU MICROMAX-007 / Wavelength: 1.54 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller






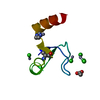
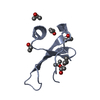




 PDBj
PDBj










