+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1m8a | ||||||
|---|---|---|---|---|---|---|---|
| Title | Human MIP-3alpha/CCL20 | ||||||
 Components Components | Small inducible cytokine A20 | ||||||
 Keywords Keywords | CYTOKINE / CC-chemokine / IL-8 type dimer | ||||||
| Function / homology |  Function and homology information Function and homology informationthymocyte migration / CCR6 chemokine receptor binding / chemokine activity / Chemokine receptors bind chemokines / Interleukin-10 signaling / positive regulation of T cell migration / T cell migration / cell chemotaxis / calcium-mediated signaling / chemotaxis ...thymocyte migration / CCR6 chemokine receptor binding / chemokine activity / Chemokine receptors bind chemokines / Interleukin-10 signaling / positive regulation of T cell migration / T cell migration / cell chemotaxis / calcium-mediated signaling / chemotaxis / cell-cell signaling / G alpha (i) signalling events / defense response to bacterium / immune response / inflammatory response / signal transduction / : / extracellular region Similarity search - Function | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | ||||||
 Authors Authors | Hoover, D.M. / Boulegue, C. / Yang, D. / Oppenheim, J.J. / Tucker, K. / Lu, W. / Lubkowski, J. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2002 Journal: J.Biol.Chem. / Year: 2002Title: The structure of human macrophage inflammatory protein-3alpha /CCL20. Linking antimicrobial and CC chemokine receptor-6-binding activities with human beta-defensins Authors: Hoover, D.M. / Boulegue, C. / Yang, D. / Oppenheim, J.J. / Tucker, K. / Lu, W. / Lubkowski, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1m8a.cif.gz 1m8a.cif.gz | 43.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1m8a.ent.gz pdb1m8a.ent.gz | 30.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1m8a.json.gz 1m8a.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/m8/1m8a https://data.pdbj.org/pub/pdb/validation_reports/m8/1m8a ftp://data.pdbj.org/pub/pdb/validation_reports/m8/1m8a ftp://data.pdbj.org/pub/pdb/validation_reports/m8/1m8a | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1eqtS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 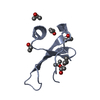
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The molecule is likely a monomer |
- Components
Components
| #1: Protein | Mass: 8043.568 Da / Num. of mol.: 2 / Source method: obtained synthetically Details: THE PROTEIN WAS SYNTHESIZED USING Machine-assisted stepwise solid phase peptide synthesis on an Applied Biosystems 433A synthesizer using a custom-designed program tailored from the ...Details: THE PROTEIN WAS SYNTHESIZED USING Machine-assisted stepwise solid phase peptide synthesis on an Applied Biosystems 433A synthesizer using a custom-designed program tailored from the published in situ neutralization/HBTU activation protocol for Boc chemistry (Schnolzer, M. et al. (1992) J.Pept.Protein Res. 40, 180-193). IT IS NATURALLY FOUND IN HOMO SAPIENS (HUMAN). References: UniProt: P78556 #2: Chemical | ChemComp-IPA / #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.69 Å3/Da / Density % sol: 54.3 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 285 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: isopropanol, sodium citrate, N-(2-hydroxyethyl)piperazine-N'-(2-ethanesulfonic acid), pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 285K | ||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 12 ℃ | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X9B / Wavelength: 0.98 Å / Beamline: X9B / Wavelength: 0.98 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Jan 25, 2002 / Details: mirrors |
| Radiation | Monochromator: si crystal / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.98 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→20 Å / Num. all: 21953 / Num. obs: 21953 / % possible obs: 97.5 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 4.5 % / Rmerge(I) obs: 0.067 / Net I/σ(I): 23.8 |
| Reflection shell | Resolution: 1.7→1.76 Å / Rmerge(I) obs: 0.623 / Mean I/σ(I) obs: 1.9 / % possible all: 96.8 |
| Reflection | *PLUS Highest resolution: 1.7 Å / Lowest resolution: 20 Å / Num. measured all: 99101 |
| Reflection shell | *PLUS Highest resolution: 1.7 Å / % possible obs: 96.8 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1EQT Resolution: 1.7→15 Å / Num. parameters: 4979 / Num. restraintsaints: 4435 Isotropic thermal model: anisotropic for sulfur, isotropic otherwise Cross valid method: FREE R / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber Details: ANISOTROPIC SCALING APPLIED BY THE METHOD OF PARKIN, MOEZZI & HOPE, J.APPL.CRYST.28(1995)53-56
| |||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: MOEWS & KRETSINGER, J.MOL.BIOL.91(1973)201-228 | |||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 27.7 Å2 | |||||||||||||||||||||||||||||||||
| Refine analyze | Num. disordered residues: 5 / Occupancy sum hydrogen: 0 / Occupancy sum non hydrogen: 1175 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→15 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||
| Software | *PLUS Name: SHELXL / Version: 97 / Classification: refinement | |||||||||||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 15 Å / % reflection Rfree: 4 % / Rfactor all: 0.19 / Rfactor Rfree: 0.238 / Rfactor Rwork: 0.19 | |||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: s_plane_restr / Dev ideal: 0.028 |
 Movie
Movie Controller
Controller












 PDBj
PDBj





