+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4nw3 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of MLL CXXC domain in complex with a CpG DNA | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA BINDING PROTEIN/DNA / Histone-lysine N-methyltransferase / CpG island / CG island / structural genomics consortium / SGC / DNA BINDING PROTEIN-DNA complex | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of DNA methylation-dependent heterochromatin formation / [histone H3]-lysine4 N-methyltransferase / histone H3K4 monomethyltransferase activity / protein-cysteine methyltransferase activity / response to potassium ion / histone H3K4 trimethyltransferase activity / unmethylated CpG binding / Phosphorylation of CLOCK, acetylation of BMAL1 (ARNTL) at target gene promoters / The CRY:PER:kinase complex represses transactivation by the BMAL:CLOCK (ARNTL:CLOCK) complex / definitive hemopoiesis ...negative regulation of DNA methylation-dependent heterochromatin formation / [histone H3]-lysine4 N-methyltransferase / histone H3K4 monomethyltransferase activity / protein-cysteine methyltransferase activity / response to potassium ion / histone H3K4 trimethyltransferase activity / unmethylated CpG binding / Phosphorylation of CLOCK, acetylation of BMAL1 (ARNTL) at target gene promoters / The CRY:PER:kinase complex represses transactivation by the BMAL:CLOCK (ARNTL:CLOCK) complex / definitive hemopoiesis / histone H3K4 methyltransferase activity / regulation of short-term neuronal synaptic plasticity / anterior/posterior pattern specification / Phosphorylated BMAL1:CLOCK (ARNTL:CLOCK) activates expression of core clock genes / embryonic hemopoiesis / T-helper 2 cell differentiation / Formation of WDR5-containing histone-modifying complexes / histone methyltransferase complex / minor groove of adenine-thymine-rich DNA binding / exploration behavior / MLL1 complex / cellular response to transforming growth factor beta stimulus / membrane depolarization / negative regulation of fibroblast proliferation / spleen development / homeostasis of number of cells within a tissue / Transferases; Transferring one-carbon groups; Methyltransferases / transcription initiation-coupled chromatin remodeling / post-embryonic development / circadian regulation of gene expression / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / visual learning / PKMTs methylate histone lysines / Transcriptional regulation of granulopoiesis / RUNX1 regulates transcription of genes involved in differentiation of HSCs / protein-containing complex assembly / fibroblast proliferation / methylation / apoptotic process / chromatin binding / positive regulation of DNA-templated transcription / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / zinc ion binding / nucleoplasm / identical protein binding / nucleus / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 2.82 Å molecular replacement / Resolution: 2.82 Å | ||||||
 Authors Authors | Bian, C. / Tempel, W. / Chao, X. / Walker, J.R. / Bountra, C. / Weigelt, J. / Arrowsmith, C.H. / Edwards, A.M. / Min, J. / Structural Genomics Consortium (SGC) | ||||||
 Citation Citation |  Journal: Structure / Year: 2018 Journal: Structure / Year: 2018Title: DNA Sequence Recognition of Human CXXC Domains and Their Structural Determinants. Authors: Xu, C. / Liu, K. / Lei, M. / Yang, A. / Li, Y. / Hughes, T.R. / Min, J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4nw3.cif.gz 4nw3.cif.gz | 61.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4nw3.ent.gz pdb4nw3.ent.gz | 42.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4nw3.json.gz 4nw3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/nw/4nw3 https://data.pdbj.org/pub/pdb/validation_reports/nw/4nw3 ftp://data.pdbj.org/pub/pdb/validation_reports/nw/4nw3 ftp://data.pdbj.org/pub/pdb/validation_reports/nw/4nw3 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4o64C  4pziC 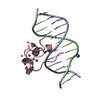 4z3cC  5vc9C  5w9qC  5w9sC  6asbC  6asdC  3qmbS  3qmgS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||||||||
| Unit cell |
| ||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS domain segments: Component-ID: _ / Ens-ID: 1 / Beg auth comp-ID: DG / Beg label comp-ID: DG / End auth comp-ID: DC / End label comp-ID: DC / Refine code: _ / Auth seq-ID: 1 - 12 / Label seq-ID: 1 - 12
|
- Components
Components
| #1: Protein | Mass: 8805.320 Da / Num. of mol.: 1 / Fragment: CXXC zinc finger domain (UNP residues 1147-1204) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: KMT2A, ALL1, CXXC7, HRX, HTRX, MLL, MLL1, TRX1 / Plasmid: pET28-mhl / Production host: Homo sapiens (human) / Gene: KMT2A, ALL1, CXXC7, HRX, HTRX, MLL, MLL1, TRX1 / Plasmid: pET28-mhl / Production host:  | ||
|---|---|---|---|
| #2: DNA chain | Mass: 3663.392 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: CpG island #3: Chemical | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.67 Å3/Da / Density % sol: 53.89 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion Details: 20% PEG3350, 0.05 M sodium tartrate, VAPOR DIFFUSION, temperature 291K |
-Data collection
| Diffraction | Mean temperature: 100 K | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 1.28231 Å / Beamline: 19-ID / Wavelength: 1.28231 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Apr 21, 2011 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: Rosenbaum-Rock high-resolution double-crystal Si(111) Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.28231 Å / Relative weight: 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.82→40 Å / Num. all: 4229 / Num. obs: 4229 / % possible obs: 99.8 % / Redundancy: 3.7 % / Rmerge(I) obs: 0.066 / Rrim(I) all: 0.066 / Χ2: 1.96 / Net I/av σ(I): 31.989 / Net I/σ(I): 12.2 / Num. measured all: 30008 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement |
|---|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRIES 3QMB AND 3QMG Resolution: 2.82→30 Å / Cor.coef. Fo:Fc: 0.957 / Cor.coef. Fo:Fc free: 0.939 / WRfactor Rfree: 0.2828 / WRfactor Rwork: 0.2543 / Occupancy max: 1 / Occupancy min: 1 / FOM work R set: 0.7519 / SU B: 34.965 / SU ML: 0.313 / SU R Cruickshank DPI: 0.7286 / SU Rfree: 0.3517 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.729 / ESU R Free: 0.352 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: AUTHORS NOTE THE SMALL NUMBER OF REFLECTIONS IN THE FREE SET, WHICH REDUCES RELIABILITY OF CROSS VALIDATION. COOT, AUTOBUSTER AND THE MOLPROBITY SERVER WERE ALSO USED DURING REFINEMENT.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 171.37 Å2 / Biso mean: 101.494 Å2 / Biso min: 67.01 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.82→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | Ens-ID: 1 / Number: 892 / Refine-ID: X-RAY DIFFRACTION / Type: interatomic distance / Rms dev position: 0.12 Å / Weight position: 0.05
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.82→2.893 Å / Total num. of bins used: 20
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj








































