+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3kiv | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
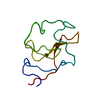
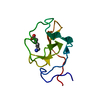
| Title | RECOMBINANT KRINGLE IV-10/M66 VARIANT OF HUMAN APOLIPOPROTEIN(A) | |||||||||
 Components Components | APOLIPOPROTEIN | |||||||||
 Keywords Keywords | KRINGLE / LYSINE BINDING SITE / APOLIPOPROTEIN(A) | |||||||||
| Function / homology |  Function and homology information Function and homology informationplasma lipoprotein particle / LDL remodeling / blood circulation / lipid transport / endopeptidase inhibitor activity / fibronectin binding / Hydrolases; Acting on peptide bonds (peptidases); Serine endopeptidases / apolipoprotein binding / lipid metabolic process / heparin binding ...plasma lipoprotein particle / LDL remodeling / blood circulation / lipid transport / endopeptidase inhibitor activity / fibronectin binding / Hydrolases; Acting on peptide bonds (peptidases); Serine endopeptidases / apolipoprotein binding / lipid metabolic process / heparin binding / serine-type endopeptidase activity / proteolysis / extracellular region Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | |||||||||
 Authors Authors | Mochalkin, I. / Tulinsky, A. / Scanu, A. | |||||||||
 Citation Citation |  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: Recombinant kringle IV-10 modules of human apolipoprotein(a): structure, ligand binding modes, and biological relevance. Authors: Mochalkin, I. / Cheng, B. / Klezovitch, O. / Scanu, A.M. / Tulinsky, A. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3kiv.cif.gz 3kiv.cif.gz | 32.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3kiv.ent.gz pdb3kiv.ent.gz | 20.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3kiv.json.gz 3kiv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ki/3kiv https://data.pdbj.org/pub/pdb/validation_reports/ki/3kiv ftp://data.pdbj.org/pub/pdb/validation_reports/ki/3kiv ftp://data.pdbj.org/pub/pdb/validation_reports/ki/3kiv | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9169.012 Da / Num. of mol.: 1 / Fragment: KRINGLE IV-10 Source method: isolated from a genetically manipulated source Details: KIV-10/T66 VARIANT OF HUMAN APOLIPOPROTEIN(A) / Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  |
|---|---|
| #2: Chemical | ChemComp-ACA / |
| #3: Water | ChemComp-HOH / |
| Compound details | N-TERMINAL (VR) AND C-TERMINAL (SDTEGTV) INTERKRINGULAR PARTS ARE DISORDERED. THE P30 RESIDUE IS IN ...N-TERMINAL (VR) AND C-TERMINAL (SDTEGTV) INTERKRING |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.76 Å3/Da / Density % sol: 30.6 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 7 / Details: pH 7.0 | ||||||||||||||||||||||||||||||||||||
| Crystal | *PLUS | ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 130 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 |
| Detector | Type: RIGAKU / Detector: IMAGE PLATE / Date: Oct 1, 1997 / Details: MSC-YALE MIRRORS |
| Radiation | Monochromator: NI FILTER / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→23 Å / Num. obs: 5656 / % possible obs: 81 % / Redundancy: 2.5 % / Rmerge(I) obs: 0.06 / Net I/σ(I): 12.8 |
| Reflection shell | Resolution: 1.8→2 Å / Rmerge(I) obs: 0.1 / Mean I/σ(I) obs: 4.7 / % possible all: 53 |
| Reflection | *PLUS Num. measured all: 12758 / Rmerge(I) obs: 0.06 |
| Reflection shell | *PLUS Highest resolution: 1.8 Å / Lowest resolution: 2 Å / % possible obs: 53 % / Rmerge(I) obs: 0.101 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.8→9 Å / σ(F): 1 MOLECULAR REPLACEMENT / Resolution: 1.8→9 Å / σ(F): 1
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 15.9 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→9 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: PROFFT / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.182 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller









 PDBj
PDBj