[English] 日本語
 Yorodumi
Yorodumi- PDB-2xdc: Structure of linear gramicidin D obtained using Type I crystals g... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2xdc | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
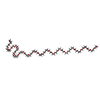
| Title | Structure of linear gramicidin D obtained using Type I crystals grown in a lipid cubic phase. | |||||||||
 Components Components | GRAMICIDIN A | |||||||||
 Keywords Keywords | ANTIBIOTIC / ION CHANNEL / MONOOLEIN / LIPID CUBIC PHASE / MESOPHASE SPONGE PHASE / BILAYER / ANTIFUNGAL / ANTIBACTERIAL | |||||||||
| Function / homology | GRAMICIDIN A / :  Function and homology information Function and homology information | |||||||||
| Biological species |  BREVIBACILLUS BREVIS (bacteria) BREVIBACILLUS BREVIS (bacteria) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | |||||||||
 Authors Authors | Hoefer, N. / Aragao, D. / Caffrey, M. | |||||||||
 Citation Citation |  Journal: Biophys.J. / Year: 2010 Journal: Biophys.J. / Year: 2010Title: Crystallizing Transmembrane Peptides in Lipidic Mesophases Authors: Hoefer, N. / Aragao, D. / Caffrey, M. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2xdc.cif.gz 2xdc.cif.gz | 41.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2xdc.ent.gz pdb2xdc.ent.gz | 29.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2xdc.json.gz 2xdc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xd/2xdc https://data.pdbj.org/pub/pdb/validation_reports/xd/2xdc ftp://data.pdbj.org/pub/pdb/validation_reports/xd/2xdc ftp://data.pdbj.org/pub/pdb/validation_reports/xd/2xdc | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1al4S S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||||||
| 2 | 
| ||||||||||||
| 3 | 
| ||||||||||||
| Unit cell |
| ||||||||||||
| Noncrystallographic symmetry (NCS) | NCS oper:
|
- Components
Components
| #1: Protein/peptide |   Type: Polypeptide / Class: Antibiotic / Mass: 1882.294 Da / Num. of mol.: 6 / Source method: isolated from a natural source Type: Polypeptide / Class: Antibiotic / Mass: 1882.294 Da / Num. of mol.: 6 / Source method: isolated from a natural sourceDetails: GRAMICIDIN A IS A HEXADECAMERIC HELICAL PEPTIDE WITH ALTERNATING D,L CHARACTERISTICS. THE N-TERM IS FORMYLATED (RESIDUE 0). THE C-TERM IS CAPPED WITH ETHANOLAMINE (RESIDUE 16). Source: (natural)  BREVIBACILLUS BREVIS (bacteria) / References: NOR: NOR00243, GRAMICIDIN A BREVIBACILLUS BREVIS (bacteria) / References: NOR: NOR00243, GRAMICIDIN A#2: Chemical | ChemComp-15P / #3: Chemical | ChemComp-NA / | #4: Water | ChemComp-HOH / | Compound details | GRAMICIDIN IS A HETEROGENEOUS MIXTURE OF SEVERAL COMPOUNDS INCLUDING GRAMICIDIN A, B AND C WHICH ...GRAMICIDIN | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.53 Å3/Da / Density % sol: 51 % Description: DATA WERE COLLECTED USING A COLLIMATED MINIBEAM WITH A 10 MICRON BEAMSIZE. |
|---|---|
| Crystal grow | Method: lipidic cubic phase / pH: 9 Details: 20 %(W/V) POLYETHYLENE GLYCOL (PEG) 6000, 0.1 M BICINE, PH 9.0. LIPIDIC CUBIC PHASE OF MONOOLEIN WAS USED IN A RATIO OF 1:20 (MOL/MOL) GRAMICIDIN D TO MONOOLEIN |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 23-ID-B / Wavelength: 1.03312 / Beamline: 23-ID-B / Wavelength: 1.03312 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Oct 16, 2008 / Details: SI(111) DOUBLE CRYSTAL |
| Radiation | Monochromator: DOUBLE CRYSTAL CRYO-COOLED / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.03312 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→31.3 Å / Num. obs: 11779 / % possible obs: 94.2 % / Observed criterion σ(I): 2 / Redundancy: 3 % / Rmerge(I) obs: 0.07 / Net I/σ(I): 13.1 |
| Reflection shell | Resolution: 1.7→1.73 Å / Redundancy: 2.2 % / Rmerge(I) obs: 0.26 / Mean I/σ(I) obs: 2 / % possible all: 80.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1AL4 Resolution: 1.7→27.15 Å / Cor.coef. Fo:Fc: 0.96 / Cor.coef. Fo:Fc free: 0.95 / SU B: 1.796 / SU ML: 0.059 / Cross valid method: THROUGHOUT / ESU R: 0.112 / ESU R Free: 0.108 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 15.243 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→27.15 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller

















 PDBj
PDBj