+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2pbn | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the human tyrosine receptor phosphate gamma | ||||||
 Components Components | Receptor-type tyrosine-protein phosphatase gamma | ||||||
 Keywords Keywords | HYDROLASE / STRUCTURAL GENOMICS / PSI-2 / Protein Structure Initiative / New York SGX Research Center for Structural Genomics / NYSGXRC | ||||||
| Function / homology |  Function and homology information Function and homology informationtransmembrane receptor protein tyrosine phosphatase activity / negative regulation of epithelial cell migration / protein-tyrosine-phosphatase / protein tyrosine phosphatase activity / cell surface receptor protein tyrosine kinase signaling pathway / neuron projection development / negative regulation of neuron projection development / signal transduction / extracellular exosome / identical protein binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | ||||||
 Authors Authors | Bonanno, J.B. / Freeman, J. / Bain, K.T. / Reyes, C. / Pelletier, L. / Jin, X. / Smith, D. / Wasserman, S. / Sauder, J.M. / Burley, S.K. ...Bonanno, J.B. / Freeman, J. / Bain, K.T. / Reyes, C. / Pelletier, L. / Jin, X. / Smith, D. / Wasserman, S. / Sauder, J.M. / Burley, S.K. / Almo, S.C. / New York SGX Research Center for Structural Genomics (NYSGXRC) | ||||||
 Citation Citation |  Journal: J.Struct.Funct.Genom. / Year: 2007 Journal: J.Struct.Funct.Genom. / Year: 2007Title: Structural genomics of protein phosphatases. Authors: Almo, S.C. / Bonanno, J.B. / Sauder, J.M. / Emtage, S. / Dilorenzo, T.P. / Malashkevich, V. / Wasserman, S.R. / Swaminathan, S. / Eswaramoorthy, S. / Agarwal, R. / Kumaran, D. / Madegowda, M. ...Authors: Almo, S.C. / Bonanno, J.B. / Sauder, J.M. / Emtage, S. / Dilorenzo, T.P. / Malashkevich, V. / Wasserman, S.R. / Swaminathan, S. / Eswaramoorthy, S. / Agarwal, R. / Kumaran, D. / Madegowda, M. / Ragumani, S. / Patskovsky, Y. / Alvarado, J. / Ramagopal, U.A. / Faber-Barata, J. / Chance, M.R. / Sali, A. / Fiser, A. / Zhang, Z.Y. / Lawrence, D.S. / Burley, S.K. | ||||||
| History |
| ||||||
| Remark 300 | BIOMOLECULE: 1 THIS ENTRY CONTAINS THE CRYSTALLOGRAPHIC ASYMMETRIC UNIT WHICH CONSISTS OF 1 ... BIOMOLECULE: 1 THIS ENTRY CONTAINS THE CRYSTALLOGRAPHIC ASYMMETRIC UNIT WHICH CONSISTS OF 1 CHAIN(S). AUTHORS STATE THAT A MONOMER IS PROBABLY THE BIOLOGICAL UNIT OF THIS POLYPEPTIDE. SEE REMARK 350 FOR INFORMATION ON GENERATING THE BIOLOGICAL MOLECULE(S). |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2pbn.cif.gz 2pbn.cif.gz | 78.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2pbn.ent.gz pdb2pbn.ent.gz | 57.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2pbn.json.gz 2pbn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pb/2pbn https://data.pdbj.org/pub/pdb/validation_reports/pb/2pbn ftp://data.pdbj.org/pub/pdb/validation_reports/pb/2pbn ftp://data.pdbj.org/pub/pdb/validation_reports/pb/2pbn | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 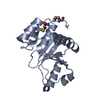 1rxdC  2fh7C  2g59C  2hcmC  2hhlC  2hxpC  2hy3SC  2i0oC  2i1yC  2i44C  2iq1C  2irmC  2isnC  2nv5C  2oycC  2p27C  2p4uC  2p69C  2p8eC  2q5eC  2qjcC  2r0bC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Details | probable monomer |
- Components
Components
| #1: Protein | Mass: 36088.941 Da / Num. of mol.: 1 / Fragment: Tyrosine-protein phosphatase 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PTPRG / Plasmid: modified pET26 / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Gene: PTPRG / Plasmid: modified pET26 / Species (production host): Escherichia coli / Production host:  |
|---|---|
| #2: Chemical | ChemComp-SO4 / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.12 Å3/Da / Density % sol: 60.59 % |
|---|---|
| Crystal grow | Temperature: 294 K / Method: vapor diffusion / pH: 6.5 Details: 100mM Sodium MES pH 6.5, 22% PEG 8000, 200mM Ammonium sulfate, VAPOR DIFFUSION, temperature 294K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X29A / Wavelength: 1.1 Å / Beamline: X29A / Wavelength: 1.1 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Sep 11, 2006 |
| Radiation | Monochromator: Si / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→53.683 Å / Num. all: 49677 / Num. obs: 49677 / % possible obs: 99.8 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 10.2 % / Biso Wilson estimate: 31.3 Å2 / Rmerge(I) obs: 0.054 / Rsym value: 0.054 / Net I/σ(I): 16.1 |
| Reflection shell | Resolution: 1.7→1.79 Å / Redundancy: 7.3 % / Rmerge(I) obs: 0.645 / Mean I/σ(I) obs: 2.2 / Num. unique all: 7108 / Rsym value: 0.645 / % possible all: 99 |
-Phasing
| Phasing MR | Rfactor: 0.403 / Cor.coef. Fo:Fc: 0.629 / Cor.coef. Io to Ic: 0.586
|
|---|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 2HY3 Resolution: 1.7→20 Å / Cor.coef. Fo:Fc: 0.961 / Cor.coef. Fo:Fc free: 0.953 / SU B: 2.158 / SU ML: 0.07 / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / ESU R: 0.095 / ESU R Free: 0.095 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 41.844 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.7→1.744 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller













 PDBj
PDBj








