+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2oyc | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of human pyridoxal phosphate phosphatase | ||||||
 Components Components | Pyridoxal phosphate phosphatase | ||||||
 Keywords Keywords | HYDROLASE / Phosphatase / Structural Genomics / NYSGXRC / New York SGX Research Center for Structural Genomics / PSI-2 / Protein Structure Initiative | ||||||
| Function / homology |  Function and homology information Function and homology informationpyridoxal phosphatase / pyridoxal 5'-phosphate catabolic process / actin rod assembly / pyridoxal phosphatase activity / positive regulation of actin filament depolymerization / dephosphorylation / regulation of modification of postsynaptic structure / protein dephosphorylation / regulation of mitotic nuclear division / protein-serine/threonine phosphatase ...pyridoxal phosphatase / pyridoxal 5'-phosphate catabolic process / actin rod assembly / pyridoxal phosphatase activity / positive regulation of actin filament depolymerization / dephosphorylation / regulation of modification of postsynaptic structure / protein dephosphorylation / regulation of mitotic nuclear division / protein-serine/threonine phosphatase / cellular response to ATP / protein serine/threonine phosphatase activity / lamellipodium membrane / phosphoprotein phosphatase activity / heat shock protein binding / regulation of cytokinesis / ruffle membrane / cell-cell junction / cytoskeleton / postsynapse / glutamatergic synapse / magnesium ion binding / protein homodimerization activity / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.72 Å MOLECULAR REPLACEMENT / Resolution: 1.72 Å | ||||||
 Authors Authors | Ramagopal, U.A. / Freeman, J. / Izuka, M. / Toro, R. / Sauder, J.M. / Burley, S.K. / Almo, S.C. / New York SGX Research Center for Structural Genomics (NYSGXRC) | ||||||
 Citation Citation |  Journal: J.Struct.Funct.Genom. / Year: 2007 Journal: J.Struct.Funct.Genom. / Year: 2007Title: Structural genomics of protein phosphatases. Authors: Almo, S.C. / Bonanno, J.B. / Sauder, J.M. / Emtage, S. / Dilorenzo, T.P. / Malashkevich, V. / Wasserman, S.R. / Swaminathan, S. / Eswaramoorthy, S. / Agarwal, R. / Kumaran, D. / Madegowda, M. ...Authors: Almo, S.C. / Bonanno, J.B. / Sauder, J.M. / Emtage, S. / Dilorenzo, T.P. / Malashkevich, V. / Wasserman, S.R. / Swaminathan, S. / Eswaramoorthy, S. / Agarwal, R. / Kumaran, D. / Madegowda, M. / Ragumani, S. / Patskovsky, Y. / Alvarado, J. / Ramagopal, U.A. / Faber-Barata, J. / Chance, M.R. / Sali, A. / Fiser, A. / Zhang, Z.Y. / Lawrence, D.S. / Burley, S.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2oyc.cif.gz 2oyc.cif.gz | 77.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2oyc.ent.gz pdb2oyc.ent.gz | 56.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2oyc.json.gz 2oyc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/oy/2oyc https://data.pdbj.org/pub/pdb/validation_reports/oy/2oyc ftp://data.pdbj.org/pub/pdb/validation_reports/oy/2oyc ftp://data.pdbj.org/pub/pdb/validation_reports/oy/2oyc | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 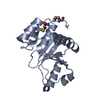 1rxdC  2fh7C  2g59C  2hcmC  2hhlC  2hxpC  2hy3C  2i0oC  2i1yC  2i44C  2iq1C  2irmC  2isnC  2nv5C  2p27C  2p4uC  2p69C  2p8eC  2pbnC  2q5eC  2qjcC  2r0bC  1zjjS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
| ||||||||
| Details | Probable dimer |
- Components
Components
| #1: Protein | Mass: 33140.996 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PDXP, PLP, PLPP / Plasmid: BC-pSGX3(BC) / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Gene: PDXP, PLP, PLPP / Plasmid: BC-pSGX3(BC) / Species (production host): Escherichia coli / Production host:  |
|---|---|
| #2: Chemical | ChemComp-NA / |
| #3: Chemical | ChemComp-WO4 / |
| #4: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.37 Å3/Da / Density % sol: 48.15 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 5.6 Details: 40% PEG4000, 0.1M Sodium citrate pH 5.6, 20% Isopropanol, VAPOR DIFFUSION, SITTING DROP, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 31-ID / Wavelength: 0.979 Å / Beamline: 31-ID / Wavelength: 0.979 Å |
| Detector | Type: MAR CCD 165 mm / Detector: CCD / Date: Feb 7, 2007 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.979 Å / Relative weight: 1 |
| Reflection | Resolution: 1.72→50 Å / Num. all: 35004 / Num. obs: 35004 / % possible obs: 99.8 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 6.6 % / Rmerge(I) obs: 0.069 / Rsym value: 0.058 / Χ2: 0.816 / Net I/σ(I): 26.4 |
| Reflection shell | Resolution: 1.72→1.78 Å / Redundancy: 6 % / Rmerge(I) obs: 0.597 / Mean I/σ(I) obs: 2.16 / Num. unique all: 6442 / Rsym value: 0.633 / Χ2: 0.514 / % possible all: 99.4 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 1ZJJ Resolution: 1.72→26.9 Å / Cor.coef. Fo:Fc: 0.962 / Cor.coef. Fo:Fc free: 0.947 / SU B: 2.36 / SU ML: 0.077 / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / ESU R: 0.111 / ESU R Free: 0.107 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 31.002 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.72→26.9 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.72→1.767 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller











 PDBj
PDBj




