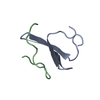
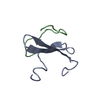
Entry Database : PDB / ID : 2ltvTitle YAP WW2 in complex with a Smad7 derived peptide Smad7 derived peptide Yorkie homolog Keywords / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / Authors Macias, M.J. / Aragon, E. / Goerner, N. / Xi, Q. / Lopes, T. / Gao, S. / Massague, J. Journal : Structure / Year : 2012Title : Structural Basis for the Versatile Interactions of Smad7 with Regulator WW Domains in TGF-beta Pathways.Authors : Aragon, E. / Goerner, N. / Xi, Q. / Gomes, T. / Gao, S. / Massague, J. / Macias, M.J. History Deposition Jun 4, 2012 Deposition site / Processing site Revision 1.0 Nov 21, 2012 Provider / Type Revision 1.1 May 1, 2024 Group / Database referencesCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_nmr_software / pdbx_nmr_spectrometer Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_nmr_software.name / _pdbx_nmr_spectrometer.model
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) Authors
Authors Citation
Citation Journal: Structure / Year: 2012
Journal: Structure / Year: 2012 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2ltv.cif.gz
2ltv.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2ltv.ent.gz
pdb2ltv.ent.gz PDB format
PDB format 2ltv.json.gz
2ltv.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/lt/2ltv
https://data.pdbj.org/pub/pdb/validation_reports/lt/2ltv ftp://data.pdbj.org/pub/pdb/validation_reports/lt/2ltv
ftp://data.pdbj.org/pub/pdb/validation_reports/lt/2ltv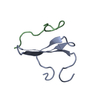

 Links
Links Assembly
Assembly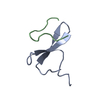
 Components
Components Homo sapiens (human) / Gene: YAP1, YAP65 / Plasmid: petM11 / Production host:
Homo sapiens (human) / Gene: YAP1, YAP65 / Plasmid: petM11 / Production host: 
 Homo sapiens (human) / References: UniProt: O15105
Homo sapiens (human) / References: UniProt: O15105 Sample preparation
Sample preparation Processing
Processing Movie
Movie Controller
Controller













 PDBj
PDBj













 HSQC
HSQC