[English] 日本語
 Yorodumi
Yorodumi- PDB-1hcj: Photoproduct of the wild-type Aequorea victoria Green Fluorescent... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1hcj | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
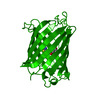
| Title | Photoproduct of the wild-type Aequorea victoria Green Fluorescent Protein | |||||||||
 Components Components | GREEN FLUORESCENT PROTEIN | |||||||||
 Keywords Keywords | LUMINESCENT PROTEIN / FLUORESCENT PROTEIN / BETA-BARREL / BIOLUMINESCENCE / LUMINESCENCE | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | |||||||||
 Authors Authors | Van Thor, J.J. / Gensch, T. / Hellingwerf, K.J. / Johnson, L. | |||||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2002 Journal: Nat.Struct.Biol. / Year: 2002Title: Phototransformation of Green Fluorescent Protein with Uv and Visible Light Leads to Decarboxylation of Glutamate 222 Authors: Van Thor, J.J. / Gensch, T. / Hellingwerf, K.J. / Johnson, L. | |||||||||
| History |
| |||||||||
| Remark 700 | SHEET DETERMINATION METHOD: DSSP THE SHEET PRESENTED FOR EACH CHAIN ON SHEET RECORDS BELOW IS ... SHEET DETERMINATION METHOD: DSSP THE SHEET PRESENTED FOR EACH CHAIN ON SHEET RECORDS BELOW IS ACTUALLY AN 11-STRANDED BARREL THAT IS REPRESENTED BY A 12-STRANDED SHEET IN WHICH THE FIRST AND LAST STRANDS ARE IDENTICAL. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1hcj.cif.gz 1hcj.cif.gz | 204.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1hcj.ent.gz pdb1hcj.ent.gz | 163.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1hcj.json.gz 1hcj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/hc/1hcj https://data.pdbj.org/pub/pdb/validation_reports/hc/1hcj ftp://data.pdbj.org/pub/pdb/validation_reports/hc/1hcj ftp://data.pdbj.org/pub/pdb/validation_reports/hc/1hcj | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1gflS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 26887.346 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Water | ChemComp-HOH / | Compound details | MUTATION GLN80ARG OTHER_DETAILS: THE CHROMOPHORE, P-HYDROXYBENZYLIDENE-IMIDAZOLIDINONE (GYS), IS ...MUTATION GLN80ARG OTHER_DETAILS: THE CHROMOPHOR | Sequence details | MODRES: 1HCJ GYS A 66() GLU 222 SIDECHAIN IS SPECIFICALLY DECARBOXYLATED AS A RESULT OF ...MODRES: 1HCJ GYS A 66() GLU 222 SIDECHAIN IS SPECIFICAL | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.34 Å3/Da / Density % sol: 43 % | |||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 277 K / pH: 7.8 Details: CRYSTALS WERE GROWN AT 4C FROM 50 MM MGCL2, 14-17 % PEG3350 AND 50-100 MM TRIS/CL PH 7.8 - 8.6. | |||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ / Method: vapor diffusion | |||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-1 / Wavelength: 0.934 / Beamline: ID14-1 / Wavelength: 0.934 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Sep 15, 2000 / Details: SAGITALLY FOCUSING GE(220 |
| Radiation | Monochromator: DIAMOND (111), GE(220) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.934 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→33.15 Å / Num. obs: 92677 / % possible obs: 97.6 % / Observed criterion σ(I): 1.5 / Redundancy: 1.9 % / Biso Wilson estimate: 23 Å2 / Rmerge(I) obs: 0.057 / Net I/σ(I): 4 |
| Reflection shell | Resolution: 1.8→1.9 Å / Redundancy: 1.9 % / Rmerge(I) obs: 0.251 / Mean I/σ(I) obs: 2.7 / % possible all: 96.8 |
| Reflection | *PLUS Num. measured all: 519063 |
| Reflection shell | *PLUS % possible obs: 96.8 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1GFL Resolution: 1.8→33.15 Å / SU B: 3.82 / SU ML: 0.118 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.149 / ESU R Free: 0.151
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 32.8 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→33.15 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: REFMAC / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.209 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller




























 PDBj
PDBj


