+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1emm | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | GREEN FLUORESCENT PROTEIN FROM AEQUOREA VICTORIA, MUTANT | ||||||||||||
 Components Components | GREEN FLUORESCENT PROTEIN | ||||||||||||
 Keywords Keywords | LUMINESCENCE / FLUORESCENT PROTEIN / BETA-BARREL / BIOLUMINESCENCE | ||||||||||||
| Function / homology |  Function and homology information Function and homology information | ||||||||||||
| Biological species |  | ||||||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||||||||
 Authors Authors | Palm, G. / Zdanov, A. / Wlodawer, A. | ||||||||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1997 Journal: Nat.Struct.Biol. / Year: 1997Title: The structural basis for spectral variations in green fluorescent protein. Authors: Palm, G.J. / Zdanov, A. / Gaitanaris, G.A. / Stauber, R. / Pavlakis, G.N. / Wlodawer, A. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1emm.cif.gz 1emm.cif.gz | 60.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1emm.ent.gz pdb1emm.ent.gz | 43.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1emm.json.gz 1emm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/em/1emm https://data.pdbj.org/pub/pdb/validation_reports/em/1emm ftp://data.pdbj.org/pub/pdb/validation_reports/em/1emm ftp://data.pdbj.org/pub/pdb/validation_reports/em/1emm | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1emcC  1emeC  1emfC  1emkC  1emlC  2emdC  2emnC  2emoC  1emaS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 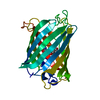
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 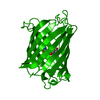
| ||||||||
| Unit cell |
| ||||||||
| Details | THE BIOLOGICALLY ACTIVE MOLECULE IS A DIMER. |
- Components
Components
| #1: Protein | Mass: 26968.416 Da / Num. of mol.: 1 / Mutation: INS(A1[B]), F64L Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.8 Å3/Da / Density % sol: 57 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Method: vapor diffusion, hanging drop / pH: 8.5 Details: PROTEIN WAS CRYSTALLIZED BY HANGING DROP METHOD. PROTEIN SOLUTION: 13 MG/ML IN 20 MM K-PO4, PH 7.0 WELL SOLUTION: 1.95 M AS, 100 MM TRIS/HCL, PH 8.5 PROTEIN:WELL 1:1, vapor diffusion - hanging drop PH range: 7.0-8.5 | |||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging dropDetails: protein solution is mixed in a 1:1 ratio with well solution | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 295 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RUH2R / Wavelength: 1.5418 |
| Detector | Type: MAC Science DIP-2000 / Detector: IMAGE PLATE / Date: Jul 25, 1996 / Details: MIRROR |
| Radiation | Monochromator: NI / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→20 Å / Num. obs: 12098 / % possible obs: 87.1 % / Observed criterion σ(I): 1 / Redundancy: 4.1 % / Rsym value: 0.07 |
| Reflection shell | Resolution: 2.3→2.4 Å / Rsym value: 0.351 / % possible all: 71.2 |
| Reflection | *PLUS Rmerge(I) obs: 0.07 |
| Reflection shell | *PLUS % possible obs: 71.2 % / Rmerge(I) obs: 0.351 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1EMA Resolution: 2.3→10 Å / Rfactor Rfree error: 0.0096 / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2 Details: PARAMETERS FOR THE CHROMOPHORE WERE ESTIMATED ACCORDING TO A MODEL COMPOUND (B.TINANT ET AL., CRYST. STRUCT. COMM., 1980, 9, 671-674)
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 33.7 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.3→2.4 Å / Rfactor Rfree error: 0.047 / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor obs: 0.2835 |
 Movie
Movie Controller
Controller












 PDBj
PDBj


