| Software | | Name | Classification |
|---|
| CNS | refinement| DENZO | data reduction| SCALEPACK | data scaling| CNS | phasing | | | |
|
|---|
| Refinement | Resolution: 2.25→45.66 Å / Rfactor Rfree error: 0.005 / Data cutoff high absF: 2893541.94 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber
| Rfactor | Num. reflection | % reflection | Selection details |
|---|
| Rfree | 0.282 | 3362 | 9.6 % | RANDOM |
|---|
| Rwork | 0.233 | - | - | - |
|---|
| all | - | 34854 | - | - |
|---|
| obs | - | 34854 | 93.1 % | - |
|---|
|
|---|
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 69.39 Å2 / ksol: 0.449 e/Å3 |
|---|
| Displacement parameters | Biso mean: 39.1 Å2
| Baniso -1 | Baniso -2 | Baniso -3 |
|---|
| 1- | 7.33 Å2 | 6 Å2 | 0 Å2 |
|---|
| 2- | - | 7.33 Å2 | 0 Å2 |
|---|
| 3- | - | - | -14.67 Å2 |
|---|
|
|---|
| Refine analyze | | Free | Obs |
|---|
| Luzzati coordinate error | 0.37 Å | 0.3 Å |
|---|
| Luzzati d res low | - | 5 Å |
|---|
| Luzzati sigma a | 0.32 Å | 0.3 Å |
|---|
|
|---|
| Refinement step | Cycle: LAST / Resolution: 2.25→45.66 Å
| Protein | Nucleic acid | Ligand | Solvent | Total |
|---|
| Num. atoms | 2618 | 0 | 6 | 111 | 2735 |
|---|
|
|---|
| Refine LS restraints | | Refine-ID | Type | Dev ideal | Dev ideal target |
|---|
| X-RAY DIFFRACTION | c_bond_d| 0.006 | | | X-RAY DIFFRACTION | c_angle_deg| 1 | | | X-RAY DIFFRACTION | c_dihedral_angle_d| 15.2 | | | X-RAY DIFFRACTION | c_improper_angle_d| 0.56 | | | X-RAY DIFFRACTION | c_mcbond_it| 3.03 | 1.5 | | X-RAY DIFFRACTION | c_mcangle_it| 4.34 | 2 | | X-RAY DIFFRACTION | c_scbond_it| 6.25 | 2 | | X-RAY DIFFRACTION | c_scangle_it| 7.76 | 2.5 | | | | | | | | |
|
|---|
| LS refinement shell | Resolution: 2.25→2.35 Å / Rfactor Rfree error: 0.021 / Total num. of bins used: 8
| Rfactor | Num. reflection | % reflection |
|---|
| Rfree | 0.348 | 284 | 9.2 % |
|---|
| Rwork | 0.303 | 2816 | - |
|---|
| obs | - | - | 65.8 % |
|---|
|
|---|
| Xplor file | | Refine-ID | Serial no | Param file | Topol file |
|---|
| X-RAY DIFFRACTION | 1 | PROTEIN_REP.PARAMPROTEIN.TOP| X-RAY DIFFRACTION | 2 | ION.PARAMION.TOP| X-RAY DIFFRACTION | 3 | WATER_REP.PARAM| WATER.TOP | | | | | |
|
|---|
| Software | *PLUS Name: CNS / Version: 1 / Classification: refinement |
|---|
| Refinement | *PLUS σ(F): 0 / % reflection Rfree: 9.6 % / Rfactor obs: 0.233 |
|---|
| Solvent computation | *PLUS |
|---|
| Displacement parameters | *PLUS Biso mean: 39.1 Å2 |
|---|
| Refine LS restraints | *PLUS | Refine-ID | Type | Dev ideal | Dev ideal target |
|---|
| X-RAY DIFFRACTION | c_angle_deg| 1 | | | X-RAY DIFFRACTION | c_dihedral_angle_d | | | X-RAY DIFFRACTION | c_dihedral_angle_deg| 15.2 | | | X-RAY DIFFRACTION | c_improper_angle_d | | | X-RAY DIFFRACTION | c_improper_angle_deg| 0.56 | | | X-RAY DIFFRACTION | c_mcbond_it | 1.5 | | X-RAY DIFFRACTION | c_scbond_it | 2 | | X-RAY DIFFRACTION | c_mcangle_it | 2 | | X-RAY DIFFRACTION | c_scangle_it | 2.5 | | | | | | | | | |
|
|---|
| LS refinement shell | *PLUS Rfactor Rfree: 0.348 / % reflection Rfree: 9.2 % / Rfactor Rwork: 0.303 |
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
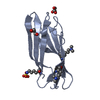
 X-RAY DIFFRACTION / Resolution: 2.25 Å
X-RAY DIFFRACTION / Resolution: 2.25 Å  Authors
Authors Citation
Citation Journal: Structure Fold.Des. / Year: 2000
Journal: Structure Fold.Des. / Year: 2000 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 1f4m.cif.gz
1f4m.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb1f4m.ent.gz
pdb1f4m.ent.gz PDB format
PDB format 1f4m.json.gz
1f4m.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/f4/1f4m
https://data.pdbj.org/pub/pdb/validation_reports/f4/1f4m ftp://data.pdbj.org/pub/pdb/validation_reports/f4/1f4m
ftp://data.pdbj.org/pub/pdb/validation_reports/f4/1f4m Links
Links Assembly
Assembly



 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU / Wavelength: 1.54128
ROTATING ANODE / Type: RIGAKU / Wavelength: 1.54128  Processing
Processing Movie
Movie Controller
Controller


















 PDBj
PDBj


