[English] 日本語
 Yorodumi
Yorodumi- PDB-1gto: HIGH RESOLUTION STRUCTURE OF A HYPERSTABLE HELICAL BUNDLE PROTEIN... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1gto | ||||||
|---|---|---|---|---|---|---|---|
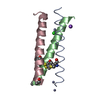
| Title | HIGH RESOLUTION STRUCTURE OF A HYPERSTABLE HELICAL BUNDLE PROTEIN MUTANT | ||||||
 Components Components | ROP | ||||||
 Keywords Keywords | TRANSCRIPTION REGULATION / TURN / HELIX PACKING / CRYSTAL CONTACTS | ||||||
| Function / homology | Helix Hairpins - #230 / Regulatory protein Rop / Rop-like superfamily / Rop protein / Helix Hairpins / Orthogonal Bundle / Mainly Alpha / identical protein binding / Regulatory protein rop Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 1.82 Å SYNCHROTRON / Resolution: 1.82 Å | ||||||
 Authors Authors | Agrawal, V. / Predki, P. / Regan, L. / Brunger, A.T. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1996 Journal: Nat.Struct.Biol. / Year: 1996Title: Amino-acid substitutions in a surface turn modulate protein stability. Authors: Predki, P.F. / Agrawal, V. / Brunger, A.T. / Regan, L. #1:  Journal: Nat.Struct.Biol. / Year: 1996 Journal: Nat.Struct.Biol. / Year: 1996Title: Amino-Acid Substitutions in a Surface Turn Modulate Protein Stability Authors: Predki, P.F. / Agrawal, V. / Brunger, A.T. / Regan, L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1gto.cif.gz 1gto.cif.gz | 46.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1gto.ent.gz pdb1gto.ent.gz | 34.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1gto.json.gz 1gto.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/gt/1gto https://data.pdbj.org/pub/pdb/validation_reports/gt/1gto ftp://data.pdbj.org/pub/pdb/validation_reports/gt/1gto ftp://data.pdbj.org/pub/pdb/validation_reports/gt/1gto | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | THREE ROP PROTOMERS ARE IN THE ASYMMETRIC UNIT, TWO OF WHICH ARE RELATED BY A NON-CRYSTALLOGRAPHIC TWO-FOLD RELATIONSHIP FORMING A ROP DIMER (B/C). THE THIRD ROP PROTOMER (A) FORMS ANOTHER ROP DIMER THROUGH A CRYSTALLOGRAPHIC SYMMETRY MATE. |
- Components
Components
| #1: Protein | Mass: 6991.702 Da / Num. of mol.: 3 / Mutation: M1G, D30G Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Water | ChemComp-HOH / | Compound details | THE HYPERSTABLE MUTANT, ROP D30G, SHOWED LITTLE INTRAMOLECULAR CHANGE FROM WILD TYPE. THE R.M.S ...THE HYPERSTABL | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.31 Å3/Da / Density % sol: 46.68 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS pH: 6.8 / Method: unknown / Details: Banner, D.W., (1987) J. Mol. Biol., 196, 657. | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL7-1 / Wavelength: 1.08 / Beamline: BL7-1 / Wavelength: 1.08 |
|---|---|
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Feb 15, 1995 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.08 Å / Relative weight: 1 |
| Reflection | Redundancy: 9 % / Biso Wilson estimate: 28.8 Å2 / Rmerge(I) obs: 0.049 |
| Reflection | *PLUS Highest resolution: 1.82 Å / Lowest resolution: 20 Å / Num. obs: 17635 / % possible obs: 98.9 % |
| Reflection shell | *PLUS Highest resolution: 1.82 Å / Lowest resolution: 1.87 Å / % possible obs: 99.4 % / Rmerge(I) obs: 0.241 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.82→20 Å / Rfactor Rfree error: 0.008 / Cross valid method: THROUGHOUT / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 32.2 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.82→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.82→1.9 Å / Rfactor Rfree error: 0.024
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.8 / Classification: refinement X-PLOR / Version: 3.8 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.28 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller











 PDBj
PDBj
