+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8476 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | human CTP synthase 1 - mutant H355A | |||||||||
 Map data Map data | human CTP synthase 1 - mutant H355A | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationhistone H1N76/N77 asparagine deamidase activity / protein asparagine deamidase activity / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / negative regulation of chromosome condensation / Interconversion of nucleotide di- and triphosphates / CTP biosynthetic process ...histone H1N76/N77 asparagine deamidase activity / protein asparagine deamidase activity / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / negative regulation of chromosome condensation / Interconversion of nucleotide di- and triphosphates / CTP biosynthetic process / nucleobase-containing compound metabolic process / Hydrolases; Acting on carbon-nitrogen bonds, other than peptide bonds; In linear amides / negative regulation of type I interferon production / DNA repair-dependent chromatin remodeling / B cell proliferation / T cell proliferation / response to xenobiotic stimulus / chromatin / ATP binding / membrane / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 17.0 Å | |||||||||
 Authors Authors | Lynch EM / Kollman JM | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2017 Journal: Nat Struct Mol Biol / Year: 2017Title: Human CTP synthase filament structure reveals the active enzyme conformation. Authors: Eric M Lynch / Derrick R Hicks / Matthew Shepherd / James A Endrizzi / Allison Maker / Jesse M Hansen / Rachael M Barry / Zemer Gitai / Enoch P Baldwin / Justin M Kollman /   Abstract: The universally conserved enzyme CTP synthase (CTPS) forms filaments in bacteria and eukaryotes. In bacteria, polymerization inhibits CTPS activity and is required for nucleotide homeostasis. Here we ...The universally conserved enzyme CTP synthase (CTPS) forms filaments in bacteria and eukaryotes. In bacteria, polymerization inhibits CTPS activity and is required for nucleotide homeostasis. Here we show that for human CTPS, polymerization increases catalytic activity. The cryo-EM structures of bacterial and human CTPS filaments differ considerably in overall architecture and in the conformation of the CTPS protomer, explaining the divergent consequences of polymerization on activity. The structure of human CTPS filament, the first structure of the full-length human enzyme, reveals a novel active conformation. The filament structures elucidate allosteric mechanisms of assembly and regulation that rely on a conserved conformational equilibrium. The findings may provide a mechanism for increasing human CTPS activity in response to metabolic state and challenge the assumption that metabolic filaments are generally storage forms of inactive enzymes. Allosteric regulation of CTPS polymerization by ligands likely represents a fundamental mechanism underlying assembly of other metabolic filaments. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8476.map.gz emd_8476.map.gz | 5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8476-v30.xml emd-8476-v30.xml emd-8476.xml emd-8476.xml | 10.4 KB 10.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8476.png emd_8476.png | 55 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8476 http://ftp.pdbj.org/pub/emdb/structures/EMD-8476 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8476 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8476 | HTTPS FTP |
-Related structure data
| Related structure data |  8474C  8475C  8490C  8491C  8504C  8513C  5tkvC  5u03C  5u05C  5u3cC  5u6rC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8476.map.gz / Format: CCP4 / Size: 5.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8476.map.gz / Format: CCP4 / Size: 5.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
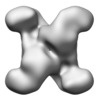
| Annotation | human CTP synthase 1 - mutant H355A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : hCTPS1-H355A tetramer
| Entire | Name: hCTPS1-H355A tetramer |
|---|---|
| Components |
|
-Supramolecule #1: hCTPS1-H355A tetramer
| Supramolecule | Name: hCTPS1-H355A tetramer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Macromolecule #1: hCTPS1-H355A
| Macromolecule | Name: hCTPS1-H355A / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MKYILVTGGV ISGIGKGIIA SSVGTILKSC GLHVTSIKID PYINIDAGTF SPYEHGEVFV LDDGGEVDL DLGNYERFLD IRLTKDNNLT TGKIYQYVIN KERKGDYLGK TVQVVPHITD A IQEWVMRQ ALIPVDEDGL EPQVCVIELG GTVGDIESMP FIEAFRQFQF ...String: MKYILVTGGV ISGIGKGIIA SSVGTILKSC GLHVTSIKID PYINIDAGTF SPYEHGEVFV LDDGGEVDL DLGNYERFLD IRLTKDNNLT TGKIYQYVIN KERKGDYLGK TVQVVPHITD A IQEWVMRQ ALIPVDEDGL EPQVCVIELG GTVGDIESMP FIEAFRQFQF KVKRENFCNI HV SLVPQPS STGEQKTKPT QNSVRELRGL GLSPDLVVCR CSNPLDTSVK EKISMFCHVE PEQ VICVHD VSSIYRVPLL LEEQGVVDYF LRRLDLPIER QPRKMLMKWK EMADRYDRLL ETCS IALVG KYTKFSDSYA SVIKALEHSA LAINHKLEIK YIDSADLEPI TSQEEPVRYA EAWQK LCSA HGVLVPGGFG VRGTEGKIQA IAWARNQKKP FLGVCLGMQL AVVEFSRNVL GWQDAN STE FDPTTSHPVV VDMPEHNPGQ MGGTMRLGKR RTLFQTKNSV MRKLYGDADY LEERHRH RF EVNPVWKKCL EEQGLKFVGQ DVEGERMEIV ELEDHPFFVG VQYHPEFLSR PIKPSPPY F GLLLASVGRL SHYLQKGCRL SPRDTYSDRS GSSSPDSEIT ELKFPSINHD |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Staining | Type: NEGATIVE / Material: Uranyl Formate |
| Details | 20 mM Tris-HCl, pH 7.9, 2 mM UTP, 2 mM ATP, 0.2 mM GTP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: CTFFIND |
|---|---|
| Final reconstruction | Applied symmetry - Point group: D2 (2x2 fold dihedral) / Resolution.type: BY AUTHOR / Resolution: 17.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 4413 |
| Initial angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION |
| Final angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)