[English] 日本語
 Yorodumi
Yorodumi- EMDB-7888: Negative Stain EM map of polyclonal serum in complex with BG505 S... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7888 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Negative Stain EM map of polyclonal serum in complex with BG505 SOSIP.664 from rabbit 3418 at post boost 2. | |||||||||
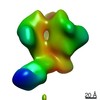
 Map data Map data | Negative stain EM map of polyclonal serum in complex with BG505 SOSIP.664 from rabbit 3418 at post boost 2 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 20.0 Å | |||||||||
 Authors Authors | Ward AB / Turner HL | |||||||||
 Citation Citation |  Journal: Immunity / Year: 2018 Journal: Immunity / Year: 2018Title: Electron-Microscopy-Based Epitope Mapping Defines Specificities of Polyclonal Antibodies Elicited during HIV-1 BG505 Envelope Trimer Immunization. Authors: Matteo Bianchi / Hannah L Turner / Bartek Nogal / Christopher A Cottrell / David Oyen / Matthias Pauthner / Raiza Bastidas / Rebecca Nedellec / Laura E McCoy / Ian A Wilson / Dennis R Burton ...Authors: Matteo Bianchi / Hannah L Turner / Bartek Nogal / Christopher A Cottrell / David Oyen / Matthias Pauthner / Raiza Bastidas / Rebecca Nedellec / Laura E McCoy / Ian A Wilson / Dennis R Burton / Andrew B Ward / Lars Hangartner /   Abstract: Characterizing polyclonal antibody responses via currently available methods is inherently complex and difficult. Mapping epitopes in an immune response is typically incomplete, which creates a ...Characterizing polyclonal antibody responses via currently available methods is inherently complex and difficult. Mapping epitopes in an immune response is typically incomplete, which creates a barrier to fully understanding the humoral response to antigens and hinders rational vaccine design efforts. Here, we describe a method of characterizing polyclonal responses by using electron microscopy, and we applied this method to the immunization of rabbits with an HIV-1 envelope glycoprotein vaccine candidate, BG505 SOSIP.664. We detected known epitopes within the polyclonal sera and revealed how antibody responses evolved during the prime-boosting strategy to ultimately result in a neutralizing antibody response. We uncovered previously unidentified epitopes, including an epitope proximal to one recognized by human broadly neutralizing antibodies as well as potentially distracting non-neutralizing epitopes. Our method provides an efficient and semiquantitative map of epitopes that are targeted in a polyclonal antibody response and should be of widespread utility in vaccine and infection studies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7888.map.gz emd_7888.map.gz | 6.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7888-v30.xml emd-7888-v30.xml emd-7888.xml emd-7888.xml | 11 KB 11 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7888.png emd_7888.png | 56.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7888 http://ftp.pdbj.org/pub/emdb/structures/EMD-7888 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7888 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7888 | HTTPS FTP |
-Related structure data
| Related structure data |  7552C  7553C  7554C  7555C  7556C  7557C  7570C  7887C  7889C  7890C  7891C  7892C  7893C  7894C  7895C  7896C  7903C  7904C  7906C  6cjkC  6didC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7888.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7888.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Negative stain EM map of polyclonal serum in complex with BG505 SOSIP.664 from rabbit 3418 at post boost 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Negative stain EM map of polyclonal serum in complex with BG505 S...
| Entire | Name: Negative stain EM map of polyclonal serum in complex with BG505 SOSIP.664 from rabbit 3418 at post boost 2. Serum digested and Fab purified before adding trimer. |
|---|---|
| Components |
|
-Supramolecule #1: Negative stain EM map of polyclonal serum in complex with BG505 S...
| Supramolecule | Name: Negative stain EM map of polyclonal serum in complex with BG505 SOSIP.664 from rabbit 3418 at post boost 2. Serum digested and Fab purified before adding trimer. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.136 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Formula: TBS |
| Staining | Type: NEGATIVE / Material: Uranyl formate |
| Grid | Model: Homemade / Material: COPPER / Mesh: 400 / Support film - Material: CELLULOSE ACETATE / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Image recording | Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 109 / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: RELION (ver. 2.1) / Number images used: 17487 |
|---|---|
| Initial angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION (ver. 2.1) |
| Final angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION (ver. 2.1) |
| Final 3D classification | Software - Name: RELION (ver. 2.1) |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















