+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4706 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bacteriophage ejection complex | ||||||||||||||||||||||||
 Map data Map data | Bacteriophage ejection complex | ||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | viral complex / DNA ejection / VIRAL PROTEIN | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationvirus tail, tube / viral portal complex / symbiont genome ejection through host cell envelope, short tail mechanism / viral DNA genome packaging Similarity search - Function | ||||||||||||||||||||||||
| Biological species |   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) | ||||||||||||||||||||||||
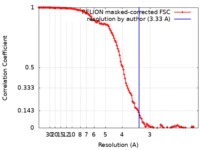
| Method | single particle reconstruction / cryo EM / Resolution: 3.33 Å | ||||||||||||||||||||||||
 Authors Authors | Cuervo A / Fabrega-Ferrer M | ||||||||||||||||||||||||
| Funding support |  Spain, 7 items Spain, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Structures of T7 bacteriophage portal and tail suggest a viral DNA retention and ejection mechanism. Authors: Ana Cuervo / Montserrat Fàbrega-Ferrer / Cristina Machón / José Javier Conesa / Francisco J Fernández / Rosa Pérez-Luque / Mar Pérez-Ruiz / Joan Pous / M Cristina Vega / José L Carrascosa / Miquel Coll /  Abstract: Double-stranded DNA bacteriophages package their genome at high pressure inside a procapsid through the portal, an oligomeric ring protein located at a unique capsid vertex. Once the DNA has been ...Double-stranded DNA bacteriophages package their genome at high pressure inside a procapsid through the portal, an oligomeric ring protein located at a unique capsid vertex. Once the DNA has been packaged, the tail components assemble on the portal to render the mature infective virion. The tail tightly seals the ejection conduit until infection, when its interaction with the host membrane triggers the opening of the channel and the viral genome is delivered to the host cell. Using high-resolution cryo-electron microscopy and X-ray crystallography, here we describe various structures of the T7 bacteriophage portal and fiber-less tail complex, which suggest a possible mechanism for DNA retention and ejection: a portal closed conformation temporarily retains the genome before the tail is assembled, whereas an open portal is found in the tail. Moreover, a fold including a seven-bladed β-propeller domain is described for the nozzle tail protein. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4706.map.gz emd_4706.map.gz | 53.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4706-v30.xml emd-4706-v30.xml emd-4706.xml emd-4706.xml | 20.3 KB 20.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4706_fsc.xml emd_4706_fsc.xml | 12.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_4706.png emd_4706.png | 1.7 MB | ||
| Filedesc metadata |  emd-4706.cif.gz emd-4706.cif.gz | 7.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4706 http://ftp.pdbj.org/pub/emdb/structures/EMD-4706 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4706 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4706 | HTTPS FTP |
-Validation report
| Summary document |  emd_4706_validation.pdf.gz emd_4706_validation.pdf.gz | 511.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4706_full_validation.pdf.gz emd_4706_full_validation.pdf.gz | 510.8 KB | Display | |
| Data in XML |  emd_4706_validation.xml.gz emd_4706_validation.xml.gz | 12.7 KB | Display | |
| Data in CIF |  emd_4706_validation.cif.gz emd_4706_validation.cif.gz | 17.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4706 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4706 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4706 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4706 | HTTPS FTP |
-Related structure data
| Related structure data |  6r21MC  4667C  4669C  6qwpC  6qx5C  6qxmC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4706.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4706.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Bacteriophage ejection complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.048 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacteriophage tail complex
| Entire | Name: Bacteriophage tail complex |
|---|---|
| Components |
|
-Supramolecule #1: Bacteriophage tail complex
| Supramolecule | Name: Bacteriophage tail complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Molecular weight | Theoretical: 534 KDa |
-Supramolecule #2: Portal protein
| Supramolecule | Name: Portal protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
-Supramolecule #3: Adaptor protein
| Supramolecule | Name: Adaptor protein / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
-Supramolecule #4: Nozzle protein
| Supramolecule | Name: Nozzle protein / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Molecular weight | Theoretical: 59.173984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAEKRTGLAE DGAKSVYERL KNDRAPYETR AQNCAQYTIP SLFPKDSDNA STDYQTPWQA VGARGLNNLA SKLMLALFPM QTWMRLTIS EYEAKQLLSD PDGLAKVDEG LSMVERIIMN YIESNSYRVT LFEALKQLVV AGNVLLYLPE PEGSNYNPMK L YRLSSYVV ...String: MAEKRTGLAE DGAKSVYERL KNDRAPYETR AQNCAQYTIP SLFPKDSDNA STDYQTPWQA VGARGLNNLA SKLMLALFPM QTWMRLTIS EYEAKQLLSD PDGLAKVDEG LSMVERIIMN YIESNSYRVT LFEALKQLVV AGNVLLYLPE PEGSNYNPMK L YRLSSYVV QRDAFGNVLQ MVTRDQIAFG ALPEDIRKAV EGQGGEKKAD ETIDVYTHIY LDEDSGEYLR YEEVEGMEVQ GS DGTYPKE ACPYIPIRMV RLDGESYGRS YIEEYLGDLR SLENLQEAIV KMSMISSKVI GLVNPAGITQ PRRLTKAQTG DFV TGRPED ISFLQLEKQA DFTVAKAVSD AIEARLSFAF MLNSAVQRTG ERVTAEEIRY VASELEDTLG GVYSILSQEL QLPL VRVLL KQLQATQQIP ELPKEAVEPT ISTGLEAIGR GQDLDKLERC VTAWAALAPM RDDPDINLAM IKLRIANAIG IDTSG ILLT EEQKQQKMAQ QSMQMGMDNG AAALAQGMAA QATASPEAMA AAADSVGLQP GI UniProtKB: Portal protein |
-Macromolecule #2: Tail tubular protein gp11
| Macromolecule | Name: Tail tubular protein gp11 / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Molecular weight | Theoretical: 26.20282 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRGSHHHHHH GMASMTGGNN MGRDLYDDDD KDPSSMRSYD MNVETAAELS AVNDILASIG EPPVSTLEGD ANADAANARR ILNKINRQI QSRGWTFNIE EGITLLPDVY SNLIVYSDDY LSLMSTSGQS IYVNRGGYVY DRTSQSDRFD SGITVNIIRL R DYDEMPEC ...String: MRGSHHHHHH GMASMTGGNN MGRDLYDDDD KDPSSMRSYD MNVETAAELS AVNDILASIG EPPVSTLEGD ANADAANARR ILNKINRQI QSRGWTFNIE EGITLLPDVY SNLIVYSDDY LSLMSTSGQS IYVNRGGYVY DRTSQSDRFD SGITVNIIRL R DYDEMPEC FRYWIVTKAS RQFNNRFFGA PEVEGVLQEE EDEARRLCME YEMDYGGYNM LDGDAFTSGL LTR UniProtKB: Tail tubular protein gp11 |
-Macromolecule #3: Tail tubular protein gp12
| Macromolecule | Name: Tail tubular protein gp12 / type: protein_or_peptide / ID: 3 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Molecular weight | Theoretical: 89.480594 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MALISQSIKN LKGGISQQPD ILRYPDQGSR QVNGWSSETE GLQKRPPLVF LNTLGDNGAL GQAPYIHLIN RDEHEQYYAV FTGSGIRVF DLSGNEKQVR YPNGSNYIKT ANPRNDLRMV TVADYTFIVN RNVVAQKNTK SVNLPNYNPN QDGLINVRGG Q YGRELIVH ...String: MALISQSIKN LKGGISQQPD ILRYPDQGSR QVNGWSSETE GLQKRPPLVF LNTLGDNGAL GQAPYIHLIN RDEHEQYYAV FTGSGIRVF DLSGNEKQVR YPNGSNYIKT ANPRNDLRMV TVADYTFIVN RNVVAQKNTK SVNLPNYNPN QDGLINVRGG Q YGRELIVH INGKDVAKYK IPDGSQPEHV NNTDAQWLAE ELAKQMRTNL SDWTVNVGQG FIHVTAPSGQ QIDSFTTKDG YA DQLINPV THYAQSFSKL PPNAPNGYMV KIVGDASKSA DQYYVRYDAE RKVWTETLGW NTEDQVLWET MPHALVRAAD GNF DFKWLE WSPKSCGDVD TNPWPSFVGS SINDVFFFRN RLGFLSGENI ILSRTAKYFN FYPASIANLS DDDPIDVAVS TNRI AILKY AVPFSEELLI WSDEAQFVLT ASGTLTSKSV ELNLTTQFDV QDRARPFGIG RNVYFASPRS SFTSIHRYYA VQDVS SVKN AEDITSHVPN YIPNGVFSIC GSGTENFCSV LSHGDPSKIF MYKFLYLNEE LRQQSWSHWD FGENVQVLAC QSISSD MYV ILRNEFNTFL ARISFTKNAI DLQGEPYRAF MDMKIRYTIP SGTYNDDTFT TSIHIPTIYG ANFGRGKITV LEPDGKI TV FEQPTAGWNS DPWLRLSGNL EGRMVYIGFN INFVYEFSKF LIKQTADDGS TSTEDIGRLQ LRRAWVNYEN SGTFDIYV E NQSSNWKYTM AGARLGSNTL RAGRLNLGTG QYRFPVVGNA KFNTVYILSD ETTPLNIIGC GWEGNYLRRS SGI UniProtKB: Tail tubular protein gp12 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 Component:
Details: 50 mM Tris-HCl pH 7.8, 100 mM NaCl, 10 mM MgCl2 | ||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER/RHODIUM / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY Details: acetone and treated with 0.1% w/v poly-L-Lysine (Sigma) for 1 min in water | ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 295.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 0.84 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-6r21: |
-Atomic model buiding 2
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Protocol: RIGID BODY FIT |
| Output model |  PDB-6r21: |
-Atomic model buiding 3
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6r21: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)