[English] 日本語
 Yorodumi
Yorodumi- EMDB-4336: Structure of two molecules of the chromatin remodelling enzyme Ch... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4336 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of two molecules of the chromatin remodelling enzyme Chd1 bound to a nucleosome | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Chromatin remodellers / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of transcriptional start site selection at RNA polymerase II promoter / nucleolar chromatin / negative regulation of DNA-templated DNA replication / regulation of chromatin organization / SLIK (SAGA-like) complex / DNA double-strand break processing / rDNA binding / nucleosome organization / nucleosome array spacer activity / histone H3K4me3 reader activity ...regulation of transcriptional start site selection at RNA polymerase II promoter / nucleolar chromatin / negative regulation of DNA-templated DNA replication / regulation of chromatin organization / SLIK (SAGA-like) complex / DNA double-strand break processing / rDNA binding / nucleosome organization / nucleosome array spacer activity / histone H3K4me3 reader activity / SAGA complex / ATP-dependent chromatin remodeler activity / sister chromatid cohesion / termination of RNA polymerase II transcription / termination of RNA polymerase I transcription / ATP-dependent activity, acting on DNA / DNA-templated transcription initiation / transcription elongation by RNA polymerase II / chromatin DNA binding / double-strand break repair via homologous recombination / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / structural constituent of chromatin / nucleosome / heterochromatin formation / nucleosome assembly / site of double-strand break / histone binding / transcription cis-regulatory region binding / chromatin remodeling / protein heterodimerization activity / chromatin binding / regulation of transcription by RNA polymerase II / chromatin / ATP hydrolysis activity / mitochondrion / DNA binding / ATP binding / nucleus Similarity search - Function | |||||||||
| Biological species |  | |||||||||
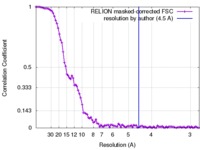
| Method | single particle reconstruction / cryo EM / Resolution: 10.0 Å | |||||||||
 Authors Authors | Sundaramoorthy R / Owen-hughes T | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2018 Journal: Elife / Year: 2018Title: Structure of the chromatin remodelling enzyme Chd1 bound to a ubiquitinylated nucleosome. Authors: Ramasubramanian Sundaramoorthy / Amanda L Hughes / Hassane El-Mkami / David G Norman / Helder Ferreira / Tom Owen-Hughes /  Abstract: ATP-dependent chromatin remodelling proteins represent a diverse family of proteins that share ATPase domains that are adapted to regulate protein-DNA interactions. Here, we present structures of the ...ATP-dependent chromatin remodelling proteins represent a diverse family of proteins that share ATPase domains that are adapted to regulate protein-DNA interactions. Here, we present structures of the Chd1 protein engaged with nucleosomes in the presence of the transition state mimic ADP-beryllium fluoride. The path of DNA strands through the ATPase domains indicates the presence of contacts conserved with single strand translocases and additional contacts with both strands that are unique to Snf2 related proteins. The structure provides connectivity between rearrangement of ATPase lobes to a closed, nucleotide bound state and the sensing of linker DNA. Two turns of linker DNA are prised off the surface of the histone octamer as a result of Chd1 binding, and both the histone H3 tail and ubiquitin conjugated to lysine 120 are re-orientated towards the unravelled DNA. This indicates how changes to nucleosome structure can alter the way in which histone epitopes are presented. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4336.map.gz emd_4336.map.gz | 55.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4336-v30.xml emd-4336-v30.xml emd-4336.xml emd-4336.xml | 26.8 KB 26.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4336_fsc.xml emd_4336_fsc.xml | 9.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_4336.png emd_4336.png | 138.5 KB | ||
| Filedesc metadata |  emd-4336.cif.gz emd-4336.cif.gz | 8.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4336 http://ftp.pdbj.org/pub/emdb/structures/EMD-4336 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4336 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4336 | HTTPS FTP |
-Related structure data
| Related structure data |  6g0lMC  4318C  6ftxC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4336.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4336.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.42 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : X. laevis nucleosome pn 601 DNA with S.cerevisiae remodeller Chd1
+Supramolecule #1: X. laevis nucleosome pn 601 DNA with S.cerevisiae remodeller Chd1
+Supramolecule #2: X. laevis nucleosome pn 601 DNA with S.cerevisiae remodeller Chd1
+Supramolecule #3: X. laevis nucleosome pn 601 DNA with S.cerevisiae remodeller Chd1
+Supramolecule #4: X. laevis nucleosome pn 601 DNA with S.cerevisiae remodeller Chd1
+Macromolecule #1: Histone H3
+Macromolecule #2: Histone H4
+Macromolecule #3: Histone H2A type 1
+Macromolecule #4: Histone H4
+Macromolecule #5: Histone H2A type 1
+Macromolecule #8: Chromo domain-containing protein 1
+Macromolecule #6: DNA (176-MER)
+Macromolecule #7: DNA (177-MER)
+Macromolecule #9: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #10: BERYLLIUM TRIFLUORIDE ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | ||||||
|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 / Component:
| ||||||
| Grid | Model: C-flat-1.2/1.3 4C / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: OTHER | ||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III | ||||||
| Details | Sample was gel filtration purified and it is monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 170.0 K / Max: 170.0 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 3870 pixel / Digitization - Dimensions - Height: 3870 pixel / Digitization - Frames/image: 5-28 / Number grids imaged: 1 / Number real images: 1800 / Average exposure time: 2.0 sec. / Average electron dose: 1.56 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 4.0 µm / Calibrated defocus min: 1.5 µm / Calibrated magnification: 98591 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 98591 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: RIGID BODY FIT / Overall B value: 800 / Target criteria: Cross-correlation coefficient |
|---|---|
| Output model |  PDB-6g0l: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)