+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1579 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Adenovirus type 5 | |||||||||
 Map data Map data | Human adenovirus type 5 at 8.9 Angstrom resolution, showing the structure of the mature, infective virion | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Human adenovirus 5 Human adenovirus 5 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.9 Å | |||||||||
 Authors Authors | Perez-Berna AJ / Marabini R / Scheres SHW / Menendez-Conejero R / Dmitriev IP / Curiel DT / Mangel WF / Flint SJ / San Martin C | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2009 Journal: J Mol Biol / Year: 2009Title: Structure and uncoating of immature adenovirus. Authors: Ana J Pérez-Berná / Roberto Marabini / Sjors H W Scheres / Rosa Menéndez-Conejero / Igor P Dmitriev / David T Curiel / Walter F Mangel / S Jane Flint / Carmen San Martín /  Abstract: Maturation via proteolytic processing is a common trait in the viral world and is often accompanied by large conformational changes and rearrangements in the capsid. The adenovirus protease has been ...Maturation via proteolytic processing is a common trait in the viral world and is often accompanied by large conformational changes and rearrangements in the capsid. The adenovirus protease has been shown to play a dual role in the viral infectious cycle: (a) in maturation, as viral assembly starts with precursors to several of the structural proteins but ends with proteolytically processed versions in the mature virion, and (b) in entry, because protease-impaired viruses have difficulties in endosome escape and uncoating. Indeed, viruses that have not undergone proteolytic processing are not infectious. We studied the three-dimensional structure of immature adenovirus particles as represented by the adenovirus type 2 thermosensitive mutant ts1 grown under non-permissive conditions and compared it with the mature capsid. Our three-dimensional electron microscopy maps at subnanometer resolution indicate that adenovirus maturation does not involve large-scale conformational changes in the capsid. Difference maps reveal the locations of unprocessed peptides pIIIa and pVI and help define their role in capsid assembly and maturation. An intriguing difference appears in the core, indicating a more compact organization and increased stability of the immature cores. We have further investigated these properties by in vitro disassembly assays. Fluorescence and electron microscopy experiments reveal differences in the stability and uncoating of immature viruses, both at the capsid and core levels, as well as disassembly intermediates not previously imaged. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1579.map.gz emd_1579.map.gz | 200.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1579-v30.xml emd-1579-v30.xml emd-1579.xml emd-1579.xml | 9.2 KB 9.2 KB | Display Display |  EMDB header EMDB header |
| Images |  1579.png 1579.png | 156.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1579 http://ftp.pdbj.org/pub/emdb/structures/EMD-1579 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1579 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1579 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1579.map.gz / Format: CCP4 / Size: 253 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1579.map.gz / Format: CCP4 / Size: 253 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human adenovirus type 5 at 8.9 Angstrom resolution, showing the structure of the mature, infective virion | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.76 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
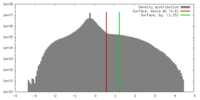
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human adenovirus type 5
| Entire | Name:   Human adenovirus type 5 Human adenovirus type 5 |
|---|---|
| Components |
|
-Supramolecule #1000: Human adenovirus type 5
| Supramolecule | Name: Human adenovirus type 5 / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Human adenovirus 5
| Supramolecule | Name: Human adenovirus 5 / type: virus / ID: 1 / Name.synonym: Adenovirus / NCBI-ID: 28285 / Sci species name: Human adenovirus 5 / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No / Syn species name: Adenovirus |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Molecular weight | Theoretical: 150 MDa |
| Virus shell | Shell ID: 1 / Diameter: 950 Å / T number (triangulation number): 25 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Details: PBS (8 mM Na2HPO4, 2 mM KH2PO4, 137 mM NaCl, 2.7 mM KCl) |
|---|---|
| Grid | Details: Quantifoil R2/4 |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM CPC / Details: Vitrification instrument: Leica CPC |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 290 / Average electron dose: 10 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.26 mm / Nominal defocus max: 5.8 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: phase flip in micrograph |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 8.9 Å / Resolution method: FSC 0.33 CUT-OFF / Software - Name: xmipp, ctffind / Number images used: 8135 |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name: URO |
| Details | PDBEntryID_givenInChain. Protocol: Rigid Body |
| Refinement | Space: RECIPROCAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)