[English] 日本語
 Yorodumi
Yorodumi- EMDB-4448: Adenovirus major core protein condenses DNA in clusters and bundl... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4448 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Adenovirus major core protein condenses DNA in clusters and bundles, modulating genome release and capsid internal pressure. | |||||||||
 Map data Map data | Human Adenovirus type 5 Ad5/attP vector with deletions in the E1 and E3 regions and an attP insertion flanking psi and a GFP cassette following psi. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Ad5/attP (virus) Ad5/attP (virus) | |||||||||
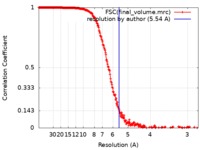
| Method | single particle reconstruction / cryo EM / Resolution: 5.54 Å | |||||||||
 Authors Authors | Hernando-Perez M / San Martin M / Condezo GN / Hearing P | |||||||||
| Funding support |  Spain, 1 items Spain, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2019 Journal: Nucleic Acids Res / Year: 2019Title: Adenovirus major core protein condenses DNA in clusters and bundles, modulating genome release and capsid internal pressure. Authors: Natalia Martín-González / Mercedes Hernando-Pérez / Gabriela N Condezo / Marta Pérez-Illana / Antonio Šiber / David Reguera / Philomena Ostapchuk / Patrick Hearing / Carmen San Martín ...Authors: Natalia Martín-González / Mercedes Hernando-Pérez / Gabriela N Condezo / Marta Pérez-Illana / Antonio Šiber / David Reguera / Philomena Ostapchuk / Patrick Hearing / Carmen San Martín / Pedro J de Pablo /    Abstract: Some viruses package dsDNA together with large amounts of positively charged proteins, thought to help condense the genome inside the capsid with no evidence. Further, this role is not clear because ...Some viruses package dsDNA together with large amounts of positively charged proteins, thought to help condense the genome inside the capsid with no evidence. Further, this role is not clear because these viruses have typically lower packing fractions than viruses encapsidating naked dsDNA. In addition, it has recently been shown that the major adenovirus condensing protein (polypeptide VII) is dispensable for genome encapsidation. Here, we study the morphology and mechanics of adenovirus particles with (Ad5-wt) and without (Ad5-VII-) protein VII. Ad5-VII- particles are stiffer than Ad5-wt, but DNA-counterions revert this difference, indicating that VII screens repulsive DNA-DNA interactions. Consequently, its absence results in increased internal pressure. The core is slightly more ordered in the absence of VII and diffuses faster out of Ad5-VII- than Ad5-wt fractured particles. In Ad5-wt unpacked cores, dsDNA associates in bundles interspersed with VII-DNA clusters. These results indicate that protein VII condenses the adenovirus genome by combining direct clustering and promotion of bridging by other core proteins. This condensation modulates the virion internal pressure and DNA release from disrupted particles, which could be crucial to keep the genome protected inside the semi-disrupted capsid while traveling to the nuclear pore. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4448.map.gz emd_4448.map.gz | 1.4 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4448-v30.xml emd-4448-v30.xml emd-4448.xml emd-4448.xml | 13.3 KB 13.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4448_fsc.xml emd_4448_fsc.xml | 26.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_4448.png emd_4448.png | 250.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4448 http://ftp.pdbj.org/pub/emdb/structures/EMD-4448 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4448 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4448 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4448.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4448.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human Adenovirus type 5 Ad5/attP vector with deletions in the E1 and E3 regions and an attP insertion flanking psi and a GFP cassette following psi. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.37 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ad5/attP
| Entire | Name:  Ad5/attP (virus) Ad5/attP (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Ad5/attP
| Supramolecule | Name: Ad5/attP / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 129951 / Sci species name: Ad5/attP / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Host system | Organism:  Homo sapiens (human) / Recombinant cell: HEK293 Homo sapiens (human) / Recombinant cell: HEK293 |
| Molecular weight | Theoretical: 150 MDa |
| Virus shell | Shell ID: 1 / Diameter: 950.0 Å / T number (triangulation number): 25 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R2/4 / Material: COPPER/RHODIUM | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM CPC | |||||||||||||||
| Details | 0.2 mg/ml initial specimen concentration was increased by consecutively incubating the grid on 10 drops of specimen before the final blotting and plunging in liquid ethane. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 1-31 / Number grids imaged: 1 / Number real images: 4395 / Average exposure time: 1.65 sec. / Average electron dose: 53.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 3.115 µm / Calibrated defocus min: 0.374 µm / Calibrated magnification: 73000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 73000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)